User login
Certain cancers primarily result from ‘bad luck’
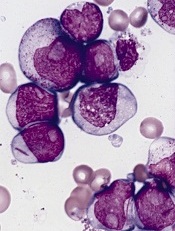
in the bone marrow
Scientists have created a statistical model that measures the proportion of cancer incidence, across many tissue types, caused mainly by random mutations that occur when stem cells divide.
By their measure, two-thirds of adult cancers—including certain leukemias—can be explained primarily by “bad luck,” when these random mutations occur in genes that can drive cancer growth.
The remaining third are due to environmental factors and inherited genes.
“All cancers are caused by a combination of bad luck, the environment, and heredity, and we’ve created a model that may help quantify how much of these three factors contribute to cancer development,” said Bert Vogelstein, MD, of the Johns Hopkins University School of Medicine.
Dr Vogelstein and Cristian Tomasetti, PhD, also of the Johns Hopkins University School of Medicine, detailed these findings in Science.
The pair came to their conclusions by searching the scientific literature for information on the cumulative number of stem cell divisions in 31 tissue types during an average individual’s lifetime.
The researchers knew that cancer arises when tissue-specific stem cells make random mistakes, or mutations. But the actual contribution of these random mistakes to cancer incidence, in comparison to the contribution of hereditary or environmental factors, was unclear.
To sort out the role of random mutations in cancer risk, the team charted the number of stem cell divisions in 31 tissues and compared these rates with the lifetime risks of cancer in the same tissues among Americans.
From this data scatterplot, Drs Tomasetti and Vogelstein determined the correlation between the total number of stem cell divisions and cancer risk to be 0.804. Mathematically, the closer this value is to 1, the more stem cell divisions and cancer risk are correlated.
“Our study shows, in general, that a change in the number of stem cell divisions in a tissue type is highly correlated with a change in the incidence of cancer in that same tissue,” Dr Vogelstein said.
One example is in colon tissue, which undergoes 4 times more stem cell divisions than small intestine tissue in humans. Likewise, colon cancer is much more prevalent than small intestinal cancer.
“You could argue that the colon is exposed to more environmental factors than the small intestine, which increases the potential rate of acquired mutations,” Dr Tomasetti said.
However, the scientists observed the opposite in mouse colons, which had a lower number of stem cell divisions than in their small intestines. In mice, cancer incidence is lower in the colon than in the small intestine. The researchers believe this supports the role of the total number of stem cell divisions in the development of cancer.
Using statistical theory, the pair calculated how much of the variation in cancer risk can be explained by the number of stem cell divisions, which is 0.804 squared, or, in percentage form, approximately 65%.
Finally, the scientists classified the types of cancers they studied into two groups. They calculated which cancer types had an incidence predicted by the number of stem cell divisions and which had higher incidence.
They found that 22 cancer types—including acute myeloid leukemia and chronic lymphocytic leukemia—could be largely explained by the “bad luck” factor of random DNA mutations during cell division.
The other 9 cancer types had incidences higher than predicted by “bad luck” and were presumably due to a combination of bad luck plus environmental or inherited factors.
“We found that the types of cancer that had higher risk than predicted by the number of stem cell divisions were precisely the ones you’d expect, including lung cancer, which is linked to smoking; skin cancer, linked to sun exposure; and forms of cancers associated with hereditary syndromes,” Dr Vogelstein said.
“This study shows that you can add to your risk of getting cancers by smoking or other poor lifestyle factors. However, many forms of cancer are due largely to the bad luck of acquiring a mutation in a cancer driver gene regardless of lifestyle and heredity factors. The best way to eradicate these cancers will be through early detection, when they are still curable by surgery.”
The researchers noted that some cancers, such as breast and prostate cancer, were not included in the report because the team was unable to find reliable stem cell division rates in the scientific literature.
They hope other scientists will help refine their statistical model by finding more precise stem cell division rates. ![]()

in the bone marrow
Scientists have created a statistical model that measures the proportion of cancer incidence, across many tissue types, caused mainly by random mutations that occur when stem cells divide.
By their measure, two-thirds of adult cancers—including certain leukemias—can be explained primarily by “bad luck,” when these random mutations occur in genes that can drive cancer growth.
The remaining third are due to environmental factors and inherited genes.
“All cancers are caused by a combination of bad luck, the environment, and heredity, and we’ve created a model that may help quantify how much of these three factors contribute to cancer development,” said Bert Vogelstein, MD, of the Johns Hopkins University School of Medicine.
Dr Vogelstein and Cristian Tomasetti, PhD, also of the Johns Hopkins University School of Medicine, detailed these findings in Science.
The pair came to their conclusions by searching the scientific literature for information on the cumulative number of stem cell divisions in 31 tissue types during an average individual’s lifetime.
The researchers knew that cancer arises when tissue-specific stem cells make random mistakes, or mutations. But the actual contribution of these random mistakes to cancer incidence, in comparison to the contribution of hereditary or environmental factors, was unclear.
To sort out the role of random mutations in cancer risk, the team charted the number of stem cell divisions in 31 tissues and compared these rates with the lifetime risks of cancer in the same tissues among Americans.
From this data scatterplot, Drs Tomasetti and Vogelstein determined the correlation between the total number of stem cell divisions and cancer risk to be 0.804. Mathematically, the closer this value is to 1, the more stem cell divisions and cancer risk are correlated.
“Our study shows, in general, that a change in the number of stem cell divisions in a tissue type is highly correlated with a change in the incidence of cancer in that same tissue,” Dr Vogelstein said.
One example is in colon tissue, which undergoes 4 times more stem cell divisions than small intestine tissue in humans. Likewise, colon cancer is much more prevalent than small intestinal cancer.
“You could argue that the colon is exposed to more environmental factors than the small intestine, which increases the potential rate of acquired mutations,” Dr Tomasetti said.
However, the scientists observed the opposite in mouse colons, which had a lower number of stem cell divisions than in their small intestines. In mice, cancer incidence is lower in the colon than in the small intestine. The researchers believe this supports the role of the total number of stem cell divisions in the development of cancer.
Using statistical theory, the pair calculated how much of the variation in cancer risk can be explained by the number of stem cell divisions, which is 0.804 squared, or, in percentage form, approximately 65%.
Finally, the scientists classified the types of cancers they studied into two groups. They calculated which cancer types had an incidence predicted by the number of stem cell divisions and which had higher incidence.
They found that 22 cancer types—including acute myeloid leukemia and chronic lymphocytic leukemia—could be largely explained by the “bad luck” factor of random DNA mutations during cell division.
The other 9 cancer types had incidences higher than predicted by “bad luck” and were presumably due to a combination of bad luck plus environmental or inherited factors.
“We found that the types of cancer that had higher risk than predicted by the number of stem cell divisions were precisely the ones you’d expect, including lung cancer, which is linked to smoking; skin cancer, linked to sun exposure; and forms of cancers associated with hereditary syndromes,” Dr Vogelstein said.
“This study shows that you can add to your risk of getting cancers by smoking or other poor lifestyle factors. However, many forms of cancer are due largely to the bad luck of acquiring a mutation in a cancer driver gene regardless of lifestyle and heredity factors. The best way to eradicate these cancers will be through early detection, when they are still curable by surgery.”
The researchers noted that some cancers, such as breast and prostate cancer, were not included in the report because the team was unable to find reliable stem cell division rates in the scientific literature.
They hope other scientists will help refine their statistical model by finding more precise stem cell division rates. ![]()

in the bone marrow
Scientists have created a statistical model that measures the proportion of cancer incidence, across many tissue types, caused mainly by random mutations that occur when stem cells divide.
By their measure, two-thirds of adult cancers—including certain leukemias—can be explained primarily by “bad luck,” when these random mutations occur in genes that can drive cancer growth.
The remaining third are due to environmental factors and inherited genes.
“All cancers are caused by a combination of bad luck, the environment, and heredity, and we’ve created a model that may help quantify how much of these three factors contribute to cancer development,” said Bert Vogelstein, MD, of the Johns Hopkins University School of Medicine.
Dr Vogelstein and Cristian Tomasetti, PhD, also of the Johns Hopkins University School of Medicine, detailed these findings in Science.
The pair came to their conclusions by searching the scientific literature for information on the cumulative number of stem cell divisions in 31 tissue types during an average individual’s lifetime.
The researchers knew that cancer arises when tissue-specific stem cells make random mistakes, or mutations. But the actual contribution of these random mistakes to cancer incidence, in comparison to the contribution of hereditary or environmental factors, was unclear.
To sort out the role of random mutations in cancer risk, the team charted the number of stem cell divisions in 31 tissues and compared these rates with the lifetime risks of cancer in the same tissues among Americans.
From this data scatterplot, Drs Tomasetti and Vogelstein determined the correlation between the total number of stem cell divisions and cancer risk to be 0.804. Mathematically, the closer this value is to 1, the more stem cell divisions and cancer risk are correlated.
“Our study shows, in general, that a change in the number of stem cell divisions in a tissue type is highly correlated with a change in the incidence of cancer in that same tissue,” Dr Vogelstein said.
One example is in colon tissue, which undergoes 4 times more stem cell divisions than small intestine tissue in humans. Likewise, colon cancer is much more prevalent than small intestinal cancer.
“You could argue that the colon is exposed to more environmental factors than the small intestine, which increases the potential rate of acquired mutations,” Dr Tomasetti said.
However, the scientists observed the opposite in mouse colons, which had a lower number of stem cell divisions than in their small intestines. In mice, cancer incidence is lower in the colon than in the small intestine. The researchers believe this supports the role of the total number of stem cell divisions in the development of cancer.
Using statistical theory, the pair calculated how much of the variation in cancer risk can be explained by the number of stem cell divisions, which is 0.804 squared, or, in percentage form, approximately 65%.
Finally, the scientists classified the types of cancers they studied into two groups. They calculated which cancer types had an incidence predicted by the number of stem cell divisions and which had higher incidence.
They found that 22 cancer types—including acute myeloid leukemia and chronic lymphocytic leukemia—could be largely explained by the “bad luck” factor of random DNA mutations during cell division.
The other 9 cancer types had incidences higher than predicted by “bad luck” and were presumably due to a combination of bad luck plus environmental or inherited factors.
“We found that the types of cancer that had higher risk than predicted by the number of stem cell divisions were precisely the ones you’d expect, including lung cancer, which is linked to smoking; skin cancer, linked to sun exposure; and forms of cancers associated with hereditary syndromes,” Dr Vogelstein said.
“This study shows that you can add to your risk of getting cancers by smoking or other poor lifestyle factors. However, many forms of cancer are due largely to the bad luck of acquiring a mutation in a cancer driver gene regardless of lifestyle and heredity factors. The best way to eradicate these cancers will be through early detection, when they are still curable by surgery.”
The researchers noted that some cancers, such as breast and prostate cancer, were not included in the report because the team was unable to find reliable stem cell division rates in the scientific literature.
They hope other scientists will help refine their statistical model by finding more precise stem cell division rates. ![]()
FDA releases warning about IV solutions
The US Food and Drug Administration (FDA) is alerting healthcare professionals not to use Wallcur, LLC, simulated intravenous (IV) products in human or animal patients, as the products are for training purposes only.
The FDA has become aware that some Wallcur training IV products have been distributed to healthcare facilities and administered to patients.
There have been reports of serious adverse events associated with some of these products, such as Practi IV Solution Bags.
Before administering IV solutions to patients, healthcare providers should carefully check the labels to ensure that products are not training products.
Wallcur’s training products, which may bear the words “for clinical simulation,” are not to be administered to patients.
If you suspect that any Wallcur training IV products may have been administered to a patient, whether or not the incident has resulted in an adverse event, please report the incident to the FDA’s MedWatch Adverse Event Reporting Program.
The FDA said it will continue to investigate and monitor this issue. The agency is also working with the Centers for Disease Control and Prevention to inform healthcare professionals and state health departments. ![]()
The US Food and Drug Administration (FDA) is alerting healthcare professionals not to use Wallcur, LLC, simulated intravenous (IV) products in human or animal patients, as the products are for training purposes only.
The FDA has become aware that some Wallcur training IV products have been distributed to healthcare facilities and administered to patients.
There have been reports of serious adverse events associated with some of these products, such as Practi IV Solution Bags.
Before administering IV solutions to patients, healthcare providers should carefully check the labels to ensure that products are not training products.
Wallcur’s training products, which may bear the words “for clinical simulation,” are not to be administered to patients.
If you suspect that any Wallcur training IV products may have been administered to a patient, whether or not the incident has resulted in an adverse event, please report the incident to the FDA’s MedWatch Adverse Event Reporting Program.
The FDA said it will continue to investigate and monitor this issue. The agency is also working with the Centers for Disease Control and Prevention to inform healthcare professionals and state health departments. ![]()
The US Food and Drug Administration (FDA) is alerting healthcare professionals not to use Wallcur, LLC, simulated intravenous (IV) products in human or animal patients, as the products are for training purposes only.
The FDA has become aware that some Wallcur training IV products have been distributed to healthcare facilities and administered to patients.
There have been reports of serious adverse events associated with some of these products, such as Practi IV Solution Bags.
Before administering IV solutions to patients, healthcare providers should carefully check the labels to ensure that products are not training products.
Wallcur’s training products, which may bear the words “for clinical simulation,” are not to be administered to patients.
If you suspect that any Wallcur training IV products may have been administered to a patient, whether or not the incident has resulted in an adverse event, please report the incident to the FDA’s MedWatch Adverse Event Reporting Program.
The FDA said it will continue to investigate and monitor this issue. The agency is also working with the Centers for Disease Control and Prevention to inform healthcare professionals and state health departments. ![]()
Lens-free microscope a ‘milestone’
Researchers say they’ve developed a lens-free microscope that can detect the presence of cell-level abnormalities with the same accuracy as larger and more expensive optical microscopes.
The invention could lead to less expensive, more portable technology for performing examinations of tissue, blood, and other biomedical specimens, according to the researchers.
It may prove especially useful in remote areas and when large numbers of samples need to be examined quickly.
Aydogan Ozcan, PhD, of the University of California, Los Angeles, and his colleagues described their work with the microscope in Science Translational Medicine.
“This is a milestone in the work we’ve been doing,” Dr Ozcan said. “This is the first time tissue samples have been imaged in 3D using a lens-free, on-chip microscope.”
The device works by using a laser or light-emitting-diode to illuminate a tissue or blood sample that has been placed on a slide and inserted into the device. A sensor array on a microchip captures and records the pattern of shadows created by the sample.
The device processes these patterns as a series of holograms, forming 3-D images of the specimen and giving medical personnel a virtual depth-of-field view. An algorithm color codes the reconstructed images, making the contrasts in the samples more apparent than they would be in the holograms and making any abnormalities easier to detect.
Dr Ozcan’s team tested the device using blood samples containing sickle cell anemia, Pap smears that indicated cervical cancer, and tissue specimens containing cancerous breast cells.
In a blind test, a board-certified pathologist analyzed sets of specimen images that had been created by the lens-free technology and by conventional microscopes. The pathologist’s diagnoses using the lens-free microscopic images proved accurate 99% of the time.
Another benefit of the lens-free device, according to the researchers, is that it produces images that are several hundred times larger in area, or field of view, than those captured by conventional bright-field optical microscopes. This makes it possible to process specimens more quickly.
“While mobile healthcare has expanded rapidly with the growth of consumer electronics—cellphones in particular—pathology is still, by and large, constrained to advanced clinical laboratory settings,” Dr Ozcan said. “Accompanied by advances in its graphical user interface, this platform could scale up for use in clinical, biomedical, scientific, educational, and citizen-science applications, among others.” ![]()
Researchers say they’ve developed a lens-free microscope that can detect the presence of cell-level abnormalities with the same accuracy as larger and more expensive optical microscopes.
The invention could lead to less expensive, more portable technology for performing examinations of tissue, blood, and other biomedical specimens, according to the researchers.
It may prove especially useful in remote areas and when large numbers of samples need to be examined quickly.
Aydogan Ozcan, PhD, of the University of California, Los Angeles, and his colleagues described their work with the microscope in Science Translational Medicine.
“This is a milestone in the work we’ve been doing,” Dr Ozcan said. “This is the first time tissue samples have been imaged in 3D using a lens-free, on-chip microscope.”
The device works by using a laser or light-emitting-diode to illuminate a tissue or blood sample that has been placed on a slide and inserted into the device. A sensor array on a microchip captures and records the pattern of shadows created by the sample.
The device processes these patterns as a series of holograms, forming 3-D images of the specimen and giving medical personnel a virtual depth-of-field view. An algorithm color codes the reconstructed images, making the contrasts in the samples more apparent than they would be in the holograms and making any abnormalities easier to detect.
Dr Ozcan’s team tested the device using blood samples containing sickle cell anemia, Pap smears that indicated cervical cancer, and tissue specimens containing cancerous breast cells.
In a blind test, a board-certified pathologist analyzed sets of specimen images that had been created by the lens-free technology and by conventional microscopes. The pathologist’s diagnoses using the lens-free microscopic images proved accurate 99% of the time.
Another benefit of the lens-free device, according to the researchers, is that it produces images that are several hundred times larger in area, or field of view, than those captured by conventional bright-field optical microscopes. This makes it possible to process specimens more quickly.
“While mobile healthcare has expanded rapidly with the growth of consumer electronics—cellphones in particular—pathology is still, by and large, constrained to advanced clinical laboratory settings,” Dr Ozcan said. “Accompanied by advances in its graphical user interface, this platform could scale up for use in clinical, biomedical, scientific, educational, and citizen-science applications, among others.” ![]()
Researchers say they’ve developed a lens-free microscope that can detect the presence of cell-level abnormalities with the same accuracy as larger and more expensive optical microscopes.
The invention could lead to less expensive, more portable technology for performing examinations of tissue, blood, and other biomedical specimens, according to the researchers.
It may prove especially useful in remote areas and when large numbers of samples need to be examined quickly.
Aydogan Ozcan, PhD, of the University of California, Los Angeles, and his colleagues described their work with the microscope in Science Translational Medicine.
“This is a milestone in the work we’ve been doing,” Dr Ozcan said. “This is the first time tissue samples have been imaged in 3D using a lens-free, on-chip microscope.”
The device works by using a laser or light-emitting-diode to illuminate a tissue or blood sample that has been placed on a slide and inserted into the device. A sensor array on a microchip captures and records the pattern of shadows created by the sample.
The device processes these patterns as a series of holograms, forming 3-D images of the specimen and giving medical personnel a virtual depth-of-field view. An algorithm color codes the reconstructed images, making the contrasts in the samples more apparent than they would be in the holograms and making any abnormalities easier to detect.
Dr Ozcan’s team tested the device using blood samples containing sickle cell anemia, Pap smears that indicated cervical cancer, and tissue specimens containing cancerous breast cells.
In a blind test, a board-certified pathologist analyzed sets of specimen images that had been created by the lens-free technology and by conventional microscopes. The pathologist’s diagnoses using the lens-free microscopic images proved accurate 99% of the time.
Another benefit of the lens-free device, according to the researchers, is that it produces images that are several hundred times larger in area, or field of view, than those captured by conventional bright-field optical microscopes. This makes it possible to process specimens more quickly.
“While mobile healthcare has expanded rapidly with the growth of consumer electronics—cellphones in particular—pathology is still, by and large, constrained to advanced clinical laboratory settings,” Dr Ozcan said. “Accompanied by advances in its graphical user interface, this platform could scale up for use in clinical, biomedical, scientific, educational, and citizen-science applications, among others.” ![]()
How malaria parasites evade the immune system
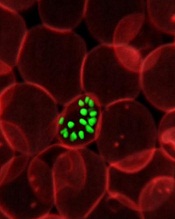
Credit: St Jude
Children’s Research Hospital
A new study has shown that malaria parasites can rapidly change proteins on the surface of human red blood cells (RBCs) during the course of a single infection, which helps the parasites evade the immune system.
The findings, which overturn previous thinking about the Plasmodium falciparum parasite’s lifecycle, could explain why so many attempts to create an effective malaria vaccine have failed and how the parasites are able to survive in the human body for such long periods of time.
Investigators described this research in PLOS Genetics.
The team kept P falciparum parasites dividing in human blood in the lab for over a year and sequenced the full parasite genome regularly. This provided snapshots of the parasite’s genome at multiple time points, allowing them to track evolution as it unfolded in the lab.
They found that the 60 or so genes that control proteins on the surface of infected human RBCs, known as var genes, swapped genetic information regularly, creating around a million new and unrecognizable surface proteins in every infected human every 2 days.
“These genes are like decks of cards constantly being shuffled,” explained study author William Hamilton, a graduate student at the Wellcome Trust Sanger Institute in Cambridge, UK.
“The use of whole-genome sequencing and the sheer number of samples we collected gave us a detailed picture of how the var gene repertoire changes continuously within red blood cells.”
The results showed, for the first time, that recombination does not occur when the malaria parasite is inside the mosquito, as previously thought. Instead, it occurs during the asexual stage of the parasite’s lifecycle inside human RBCs. This finding may help explain how chronic asymptomatic infection, a crucial problem for malaria elimination, is possible.
“It’s very likely that mosquitos are re-infected with Plasmodium falciparum parasites at the beginning of each wet season by biting humans who have carried the parasites, often asymptomatically, for up to 8 months during the dry season,” said study author Antoine Claessens, PhD, of the Wellcome Trust Sanger Institute.
“During those months, the parasite’s var genes are busy recombining to create millions of different versions—cunning disguises that mean they remain safe from the immune system and ready for the new malarial season.”
While further work will be required to fully understand the mechanism driving the recombination of P falciparum’s var genes, the investigators were able to calculate the rate at which it happens. They found that var gene recombination takes place in about 0.2% of parasites after each 48-hour life cycle in the RBC.
With about a billion parasites living inside a typical infected human, there is huge potential for the parasite to create new, recombined var genes inside each person with malaria. This pace of change far exceeds that of genes in any other region of the parasite’s genome.
“When you consider that 200 million people across the world are infected with malaria, and each of them is harboring parasites that are continually generating millions of antigenic variants, it becomes apparent why our fight against malaria is so challenging,” said study author Dominic Kwiatkowski, MBBS, of the Wellcome Trust Sanger Institute.
“This study is a great example of how genome sequence analysis is enriching our understanding of malaria biology. By learning the genetic tricks that the parasite uses to evade the human immune system, we will be in a much better position to eliminate this deadly disease.” ![]()

Credit: St Jude
Children’s Research Hospital
A new study has shown that malaria parasites can rapidly change proteins on the surface of human red blood cells (RBCs) during the course of a single infection, which helps the parasites evade the immune system.
The findings, which overturn previous thinking about the Plasmodium falciparum parasite’s lifecycle, could explain why so many attempts to create an effective malaria vaccine have failed and how the parasites are able to survive in the human body for such long periods of time.
Investigators described this research in PLOS Genetics.
The team kept P falciparum parasites dividing in human blood in the lab for over a year and sequenced the full parasite genome regularly. This provided snapshots of the parasite’s genome at multiple time points, allowing them to track evolution as it unfolded in the lab.
They found that the 60 or so genes that control proteins on the surface of infected human RBCs, known as var genes, swapped genetic information regularly, creating around a million new and unrecognizable surface proteins in every infected human every 2 days.
“These genes are like decks of cards constantly being shuffled,” explained study author William Hamilton, a graduate student at the Wellcome Trust Sanger Institute in Cambridge, UK.
“The use of whole-genome sequencing and the sheer number of samples we collected gave us a detailed picture of how the var gene repertoire changes continuously within red blood cells.”
The results showed, for the first time, that recombination does not occur when the malaria parasite is inside the mosquito, as previously thought. Instead, it occurs during the asexual stage of the parasite’s lifecycle inside human RBCs. This finding may help explain how chronic asymptomatic infection, a crucial problem for malaria elimination, is possible.
“It’s very likely that mosquitos are re-infected with Plasmodium falciparum parasites at the beginning of each wet season by biting humans who have carried the parasites, often asymptomatically, for up to 8 months during the dry season,” said study author Antoine Claessens, PhD, of the Wellcome Trust Sanger Institute.
“During those months, the parasite’s var genes are busy recombining to create millions of different versions—cunning disguises that mean they remain safe from the immune system and ready for the new malarial season.”
While further work will be required to fully understand the mechanism driving the recombination of P falciparum’s var genes, the investigators were able to calculate the rate at which it happens. They found that var gene recombination takes place in about 0.2% of parasites after each 48-hour life cycle in the RBC.
With about a billion parasites living inside a typical infected human, there is huge potential for the parasite to create new, recombined var genes inside each person with malaria. This pace of change far exceeds that of genes in any other region of the parasite’s genome.
“When you consider that 200 million people across the world are infected with malaria, and each of them is harboring parasites that are continually generating millions of antigenic variants, it becomes apparent why our fight against malaria is so challenging,” said study author Dominic Kwiatkowski, MBBS, of the Wellcome Trust Sanger Institute.
“This study is a great example of how genome sequence analysis is enriching our understanding of malaria biology. By learning the genetic tricks that the parasite uses to evade the human immune system, we will be in a much better position to eliminate this deadly disease.” ![]()

Credit: St Jude
Children’s Research Hospital
A new study has shown that malaria parasites can rapidly change proteins on the surface of human red blood cells (RBCs) during the course of a single infection, which helps the parasites evade the immune system.
The findings, which overturn previous thinking about the Plasmodium falciparum parasite’s lifecycle, could explain why so many attempts to create an effective malaria vaccine have failed and how the parasites are able to survive in the human body for such long periods of time.
Investigators described this research in PLOS Genetics.
The team kept P falciparum parasites dividing in human blood in the lab for over a year and sequenced the full parasite genome regularly. This provided snapshots of the parasite’s genome at multiple time points, allowing them to track evolution as it unfolded in the lab.
They found that the 60 or so genes that control proteins on the surface of infected human RBCs, known as var genes, swapped genetic information regularly, creating around a million new and unrecognizable surface proteins in every infected human every 2 days.
“These genes are like decks of cards constantly being shuffled,” explained study author William Hamilton, a graduate student at the Wellcome Trust Sanger Institute in Cambridge, UK.
“The use of whole-genome sequencing and the sheer number of samples we collected gave us a detailed picture of how the var gene repertoire changes continuously within red blood cells.”
The results showed, for the first time, that recombination does not occur when the malaria parasite is inside the mosquito, as previously thought. Instead, it occurs during the asexual stage of the parasite’s lifecycle inside human RBCs. This finding may help explain how chronic asymptomatic infection, a crucial problem for malaria elimination, is possible.
“It’s very likely that mosquitos are re-infected with Plasmodium falciparum parasites at the beginning of each wet season by biting humans who have carried the parasites, often asymptomatically, for up to 8 months during the dry season,” said study author Antoine Claessens, PhD, of the Wellcome Trust Sanger Institute.
“During those months, the parasite’s var genes are busy recombining to create millions of different versions—cunning disguises that mean they remain safe from the immune system and ready for the new malarial season.”
While further work will be required to fully understand the mechanism driving the recombination of P falciparum’s var genes, the investigators were able to calculate the rate at which it happens. They found that var gene recombination takes place in about 0.2% of parasites after each 48-hour life cycle in the RBC.
With about a billion parasites living inside a typical infected human, there is huge potential for the parasite to create new, recombined var genes inside each person with malaria. This pace of change far exceeds that of genes in any other region of the parasite’s genome.
“When you consider that 200 million people across the world are infected with malaria, and each of them is harboring parasites that are continually generating millions of antigenic variants, it becomes apparent why our fight against malaria is so challenging,” said study author Dominic Kwiatkowski, MBBS, of the Wellcome Trust Sanger Institute.
“This study is a great example of how genome sequence analysis is enriching our understanding of malaria biology. By learning the genetic tricks that the parasite uses to evade the human immune system, we will be in a much better position to eliminate this deadly disease.” ![]()
Obokata fails to create STAP cells, resigns

Credit: Associated Press
After months of trying, Haruko Obokata, PhD, and a team of researchers at her institution, RIKEN, have failed to produce stimulus-triggered acquisition of pluripotency (STAP) cells.
Officials from RIKEN said they have accepted Dr Obokata’s resignation, and the institution has decided to end its efforts to recreate the STAP cell phenomenon.
Dr Obokata and her colleagues initially reported the creation of STAP cells in an article and a letter published in Nature last January. The researchers said they had induced pluripotency in somatic cells by exposing the cells to a low-pH environment.
Not long after the papers were published, members of the scientific community began to question the validity of the research.
So RIKEN launched an investigation, ultimately concluding that Dr Obokata was guilty of misconduct, and some of her colleagues—including the deceased Yoshiki Sasai, MD, PhD—were guilty of negligence.
RIKEN also called for the papers to be retracted, and, in July, they were.
Throughout these proceedings, Dr Obokata insisted the STAP cell phenomenon is real. To investigate this claim, RIKEN organized a group of researchers to recreate Dr Obokata’s experiments.
In August, the group reported initial results, saying their attempts had failed, but they would continue trying to create STAP cells until March 2015. Meanwhile, Dr Obokata was trying to recreate the STAP cell phenomenon on her own, under supervision.
Shinichi Aizawa, PhD, the leader of RIKEN’s team, explained the final results of their experiments, as well as Dr Obokata’s, in a press conference in Tokyo on Friday.
Dr Obokata was able to show a fluorescent phenomenon that indicates the possibility of pluripotency in cells, albeit at a very low rate. However, she could not confirm the pluripotency of STAP cells in mice.
The RIKEN team had similar results. So they have decided not to continue with the experiments.
RIKEN accepted Dr Obokata’s resignation, and a disciplinary committee has been discussing how they will reprimand her for research misconduct. RIKEN officials said they will make an announcement once the decision has been made. ![]()

Credit: Associated Press
After months of trying, Haruko Obokata, PhD, and a team of researchers at her institution, RIKEN, have failed to produce stimulus-triggered acquisition of pluripotency (STAP) cells.
Officials from RIKEN said they have accepted Dr Obokata’s resignation, and the institution has decided to end its efforts to recreate the STAP cell phenomenon.
Dr Obokata and her colleagues initially reported the creation of STAP cells in an article and a letter published in Nature last January. The researchers said they had induced pluripotency in somatic cells by exposing the cells to a low-pH environment.
Not long after the papers were published, members of the scientific community began to question the validity of the research.
So RIKEN launched an investigation, ultimately concluding that Dr Obokata was guilty of misconduct, and some of her colleagues—including the deceased Yoshiki Sasai, MD, PhD—were guilty of negligence.
RIKEN also called for the papers to be retracted, and, in July, they were.
Throughout these proceedings, Dr Obokata insisted the STAP cell phenomenon is real. To investigate this claim, RIKEN organized a group of researchers to recreate Dr Obokata’s experiments.
In August, the group reported initial results, saying their attempts had failed, but they would continue trying to create STAP cells until March 2015. Meanwhile, Dr Obokata was trying to recreate the STAP cell phenomenon on her own, under supervision.
Shinichi Aizawa, PhD, the leader of RIKEN’s team, explained the final results of their experiments, as well as Dr Obokata’s, in a press conference in Tokyo on Friday.
Dr Obokata was able to show a fluorescent phenomenon that indicates the possibility of pluripotency in cells, albeit at a very low rate. However, she could not confirm the pluripotency of STAP cells in mice.
The RIKEN team had similar results. So they have decided not to continue with the experiments.
RIKEN accepted Dr Obokata’s resignation, and a disciplinary committee has been discussing how they will reprimand her for research misconduct. RIKEN officials said they will make an announcement once the decision has been made. ![]()

Credit: Associated Press
After months of trying, Haruko Obokata, PhD, and a team of researchers at her institution, RIKEN, have failed to produce stimulus-triggered acquisition of pluripotency (STAP) cells.
Officials from RIKEN said they have accepted Dr Obokata’s resignation, and the institution has decided to end its efforts to recreate the STAP cell phenomenon.
Dr Obokata and her colleagues initially reported the creation of STAP cells in an article and a letter published in Nature last January. The researchers said they had induced pluripotency in somatic cells by exposing the cells to a low-pH environment.
Not long after the papers were published, members of the scientific community began to question the validity of the research.
So RIKEN launched an investigation, ultimately concluding that Dr Obokata was guilty of misconduct, and some of her colleagues—including the deceased Yoshiki Sasai, MD, PhD—were guilty of negligence.
RIKEN also called for the papers to be retracted, and, in July, they were.
Throughout these proceedings, Dr Obokata insisted the STAP cell phenomenon is real. To investigate this claim, RIKEN organized a group of researchers to recreate Dr Obokata’s experiments.
In August, the group reported initial results, saying their attempts had failed, but they would continue trying to create STAP cells until March 2015. Meanwhile, Dr Obokata was trying to recreate the STAP cell phenomenon on her own, under supervision.
Shinichi Aizawa, PhD, the leader of RIKEN’s team, explained the final results of their experiments, as well as Dr Obokata’s, in a press conference in Tokyo on Friday.
Dr Obokata was able to show a fluorescent phenomenon that indicates the possibility of pluripotency in cells, albeit at a very low rate. However, she could not confirm the pluripotency of STAP cells in mice.
The RIKEN team had similar results. So they have decided not to continue with the experiments.
RIKEN accepted Dr Obokata’s resignation, and a disciplinary committee has been discussing how they will reprimand her for research misconduct. RIKEN officials said they will make an announcement once the decision has been made. ![]()
People often dismiss cancer symptoms, survey suggests

Credit: NIH
People could be putting their lives at risk by dismissing potential warning signs of cancer as less serious symptoms, according to a study published in PLOS ONE.
In a survey of about 1700 people, more than half of respondents said they had experienced at least
one red-flag cancer “alarm” symptom—such as persistent, unexplained pain or an unexplained lump—during the previous 3 months, but
only 2% of them thought cancer was a possible cause.
The survey had been sent to people aged 50 and older who were registered with 3 London general practices. The questionnaire listed 17 symptoms, including 10 widely publicized potential cancer warning signs, such as an unexplained cough, bleeding, and a persistent change in bowel or bladder habits.
Cancer was not mentioned, but the survey asked which of the symptoms subjects had experienced, what they thought caused them, if they were concerned that symptoms were serious, and whether they had consulted their doctor.
Of the 1724 subjects who responded, 53% had experienced at least one cancer “alarm” symptom in the previous 3 months.
This included unexplained cough or hoarseness; persistent change in bowel habits; persistent, unexplained pain; persistent change in bladder habits; unexplained lump; a change in the appearance of a mole; a sore that does not heal; unexplained bleeding; unexplained weight loss; and persistent difficulty swallowing.
Persistent cough (20%) and persistent change in bowel habits (18%) were the most common symptoms. Difficulty swallowing and unexplained weight loss (both 4%) were least common.
Overall, subjects appraised the cancer warning “alarm” symptoms as more serious than “non-alarm” symptoms, such as sore throat and feeling tired. Fifty-nine percent of respondents said they contacted a doctor about their “alarm” symptoms.
However, subjects rarely attributed potential signs of cancer to the disease, putting them down to other reasons, such as age, infection, arthritis, piles, and cysts.
“Most people with potential warning symptoms don’t have cancer, but some will, and others may have other diseases that would benefit from early attention,” said study author Katriina Whitaker, PhD, of University College London in the UK.
“That’s why it’s important that these symptoms are checked out, especially if they don’t go away. But people could delay seeing a doctor if they don’t acknowledge cancer as a possible cause. It’s worrying that even the more obvious warning symptoms, such as unexplained lumps or changes to the appearance of a mole, were rarely attributed to cancer, although they are often well recognized in surveys that assess the public’s knowledge of the disease.”
“Even when people thought warning symptoms might be serious, cancer didn’t tend to spring to mind. This might be because people were frightened and reluctant to mention cancer, thought cancer wouldn’t happen to them, or believed other causes were more likely.” ![]()

Credit: NIH
People could be putting their lives at risk by dismissing potential warning signs of cancer as less serious symptoms, according to a study published in PLOS ONE.
In a survey of about 1700 people, more than half of respondents said they had experienced at least
one red-flag cancer “alarm” symptom—such as persistent, unexplained pain or an unexplained lump—during the previous 3 months, but
only 2% of them thought cancer was a possible cause.
The survey had been sent to people aged 50 and older who were registered with 3 London general practices. The questionnaire listed 17 symptoms, including 10 widely publicized potential cancer warning signs, such as an unexplained cough, bleeding, and a persistent change in bowel or bladder habits.
Cancer was not mentioned, but the survey asked which of the symptoms subjects had experienced, what they thought caused them, if they were concerned that symptoms were serious, and whether they had consulted their doctor.
Of the 1724 subjects who responded, 53% had experienced at least one cancer “alarm” symptom in the previous 3 months.
This included unexplained cough or hoarseness; persistent change in bowel habits; persistent, unexplained pain; persistent change in bladder habits; unexplained lump; a change in the appearance of a mole; a sore that does not heal; unexplained bleeding; unexplained weight loss; and persistent difficulty swallowing.
Persistent cough (20%) and persistent change in bowel habits (18%) were the most common symptoms. Difficulty swallowing and unexplained weight loss (both 4%) were least common.
Overall, subjects appraised the cancer warning “alarm” symptoms as more serious than “non-alarm” symptoms, such as sore throat and feeling tired. Fifty-nine percent of respondents said they contacted a doctor about their “alarm” symptoms.
However, subjects rarely attributed potential signs of cancer to the disease, putting them down to other reasons, such as age, infection, arthritis, piles, and cysts.
“Most people with potential warning symptoms don’t have cancer, but some will, and others may have other diseases that would benefit from early attention,” said study author Katriina Whitaker, PhD, of University College London in the UK.
“That’s why it’s important that these symptoms are checked out, especially if they don’t go away. But people could delay seeing a doctor if they don’t acknowledge cancer as a possible cause. It’s worrying that even the more obvious warning symptoms, such as unexplained lumps or changes to the appearance of a mole, were rarely attributed to cancer, although they are often well recognized in surveys that assess the public’s knowledge of the disease.”
“Even when people thought warning symptoms might be serious, cancer didn’t tend to spring to mind. This might be because people were frightened and reluctant to mention cancer, thought cancer wouldn’t happen to them, or believed other causes were more likely.” ![]()

Credit: NIH
People could be putting their lives at risk by dismissing potential warning signs of cancer as less serious symptoms, according to a study published in PLOS ONE.
In a survey of about 1700 people, more than half of respondents said they had experienced at least
one red-flag cancer “alarm” symptom—such as persistent, unexplained pain or an unexplained lump—during the previous 3 months, but
only 2% of them thought cancer was a possible cause.
The survey had been sent to people aged 50 and older who were registered with 3 London general practices. The questionnaire listed 17 symptoms, including 10 widely publicized potential cancer warning signs, such as an unexplained cough, bleeding, and a persistent change in bowel or bladder habits.
Cancer was not mentioned, but the survey asked which of the symptoms subjects had experienced, what they thought caused them, if they were concerned that symptoms were serious, and whether they had consulted their doctor.
Of the 1724 subjects who responded, 53% had experienced at least one cancer “alarm” symptom in the previous 3 months.
This included unexplained cough or hoarseness; persistent change in bowel habits; persistent, unexplained pain; persistent change in bladder habits; unexplained lump; a change in the appearance of a mole; a sore that does not heal; unexplained bleeding; unexplained weight loss; and persistent difficulty swallowing.
Persistent cough (20%) and persistent change in bowel habits (18%) were the most common symptoms. Difficulty swallowing and unexplained weight loss (both 4%) were least common.
Overall, subjects appraised the cancer warning “alarm” symptoms as more serious than “non-alarm” symptoms, such as sore throat and feeling tired. Fifty-nine percent of respondents said they contacted a doctor about their “alarm” symptoms.
However, subjects rarely attributed potential signs of cancer to the disease, putting them down to other reasons, such as age, infection, arthritis, piles, and cysts.
“Most people with potential warning symptoms don’t have cancer, but some will, and others may have other diseases that would benefit from early attention,” said study author Katriina Whitaker, PhD, of University College London in the UK.
“That’s why it’s important that these symptoms are checked out, especially if they don’t go away. But people could delay seeing a doctor if they don’t acknowledge cancer as a possible cause. It’s worrying that even the more obvious warning symptoms, such as unexplained lumps or changes to the appearance of a mole, were rarely attributed to cancer, although they are often well recognized in surveys that assess the public’s knowledge of the disease.”
“Even when people thought warning symptoms might be serious, cancer didn’t tend to spring to mind. This might be because people were frightened and reluctant to mention cancer, thought cancer wouldn’t happen to them, or believed other causes were more likely.” ![]()
HHS and NIH aim to make more trial results public

Credit: Esther Dyson
The US Department of Health and Human Services (HHS) has proposed a rule that would require more public disclosure of clinical trial results.
The Notice of Proposed Rulemaking (NPRM) adds to current requirements for submitting trial information to ClinicalTrials.gov.
Most notably, the rule would require the submission of summary results from studies of products not yet approved by the US Food and Drug Administration
(FDA).
The National Institutes of Health (NIH) have created a draft policy that would extend similar reporting requirements to all clinical trials funded by NIH.
About the NPRM: Who, what, and when
The NPRM details new procedures for meeting the requirements established by the Food and Drug Administration Amendments Act of 2007 (FDAAA) to improve public access to clinical trial information.
The proposed rule specifies how information about a clinical trial would need to be submitted to ClinicalTrials.gov. It would not affect requirements for the design or conduct of clinical trials or for the data that must be collected during these trials.
The rule would apply to controlled, interventional studies of drugs, biological products, and devices that are regulated by the FDA. This excludes phase 1 studies of drugs and biological products and feasibility studies of devices.
In general, clinical trials of products regulated by the FDA will meet one or more of the following criteria: include one or more sites in the US; study a product manufactured in the US or its territories and exported for use in a trial outside the US; or be conducted under an FDA investigational new drug application or investigational device exemption.
The NPRM would require the parties responsible for applicable clinical trials—such as the sponsor or a designated principal investigator—to register the trial at ClinicalTrials.gov no later than 21 days after enrolling the first participant.
Registration consists of submitting 4 categories of data elements that are specified in the NPRM: 1) descriptive information, 2) recruitment information, 3) location and contact information, and 4) administrative information.
The parties responsible for the trial would also be required to submit a summary of the study’s results to ClinicalTrials.gov, generally no later than 12 months after trial completion.
However, the NPRM includes procedures for delaying results submission and for requesting extensions to the results submission deadline for good cause.
NIH policy: Extending the NPRM
The proposed NIH policy would make requirements in the NPRM applicable to all NIH-funded awardees and investigators conducting clinical trials, regardless of study phase, type of intervention, or whether they are subject to the rules proposed in the NPRM.
NIH awardees would be expected to ensure submission to ClinicalTrials.gov of the same type of registration and results information, and in the same timeframes, as responsible parties whose trials are subject to the FDAAA and the regulations proposed in the NPRM.
For clinical trials subject to only the proposed NIH policy (not the FDAAA), the NIH would post submitted information, in general, no later than 30 days after it is submitted.
An NIH-funded clinical trial that is also subject to FDAAA would need to have only one entry in ClinicalTrials.gov containing its registration and results information.
Open for comment
The public may comment on any aspect of the NPRM or the proposed NIH policy.
Written comments on the NPRM should be submitted to docket number NIH-2011-0003 at www.regulations.gov. Commenters should indicate the specific section of the NPRM to which each comment refers.
Written comments on the proposed NIH policy should be submitted to the Office of Clinical Research and Bioethics Policy, Office of Science Policy, NIH, via email at clinicaltrials.disseminationpolicy@mail.nih.gov, mail at 6705 Rockledge Drive, Suite 750, Bethesda, MD 20892, or fax at 301-496-9839. ![]()

Credit: Esther Dyson
The US Department of Health and Human Services (HHS) has proposed a rule that would require more public disclosure of clinical trial results.
The Notice of Proposed Rulemaking (NPRM) adds to current requirements for submitting trial information to ClinicalTrials.gov.
Most notably, the rule would require the submission of summary results from studies of products not yet approved by the US Food and Drug Administration
(FDA).
The National Institutes of Health (NIH) have created a draft policy that would extend similar reporting requirements to all clinical trials funded by NIH.
About the NPRM: Who, what, and when
The NPRM details new procedures for meeting the requirements established by the Food and Drug Administration Amendments Act of 2007 (FDAAA) to improve public access to clinical trial information.
The proposed rule specifies how information about a clinical trial would need to be submitted to ClinicalTrials.gov. It would not affect requirements for the design or conduct of clinical trials or for the data that must be collected during these trials.
The rule would apply to controlled, interventional studies of drugs, biological products, and devices that are regulated by the FDA. This excludes phase 1 studies of drugs and biological products and feasibility studies of devices.
In general, clinical trials of products regulated by the FDA will meet one or more of the following criteria: include one or more sites in the US; study a product manufactured in the US or its territories and exported for use in a trial outside the US; or be conducted under an FDA investigational new drug application or investigational device exemption.
The NPRM would require the parties responsible for applicable clinical trials—such as the sponsor or a designated principal investigator—to register the trial at ClinicalTrials.gov no later than 21 days after enrolling the first participant.
Registration consists of submitting 4 categories of data elements that are specified in the NPRM: 1) descriptive information, 2) recruitment information, 3) location and contact information, and 4) administrative information.
The parties responsible for the trial would also be required to submit a summary of the study’s results to ClinicalTrials.gov, generally no later than 12 months after trial completion.
However, the NPRM includes procedures for delaying results submission and for requesting extensions to the results submission deadline for good cause.
NIH policy: Extending the NPRM
The proposed NIH policy would make requirements in the NPRM applicable to all NIH-funded awardees and investigators conducting clinical trials, regardless of study phase, type of intervention, or whether they are subject to the rules proposed in the NPRM.
NIH awardees would be expected to ensure submission to ClinicalTrials.gov of the same type of registration and results information, and in the same timeframes, as responsible parties whose trials are subject to the FDAAA and the regulations proposed in the NPRM.
For clinical trials subject to only the proposed NIH policy (not the FDAAA), the NIH would post submitted information, in general, no later than 30 days after it is submitted.
An NIH-funded clinical trial that is also subject to FDAAA would need to have only one entry in ClinicalTrials.gov containing its registration and results information.
Open for comment
The public may comment on any aspect of the NPRM or the proposed NIH policy.
Written comments on the NPRM should be submitted to docket number NIH-2011-0003 at www.regulations.gov. Commenters should indicate the specific section of the NPRM to which each comment refers.
Written comments on the proposed NIH policy should be submitted to the Office of Clinical Research and Bioethics Policy, Office of Science Policy, NIH, via email at clinicaltrials.disseminationpolicy@mail.nih.gov, mail at 6705 Rockledge Drive, Suite 750, Bethesda, MD 20892, or fax at 301-496-9839. ![]()

Credit: Esther Dyson
The US Department of Health and Human Services (HHS) has proposed a rule that would require more public disclosure of clinical trial results.
The Notice of Proposed Rulemaking (NPRM) adds to current requirements for submitting trial information to ClinicalTrials.gov.
Most notably, the rule would require the submission of summary results from studies of products not yet approved by the US Food and Drug Administration
(FDA).
The National Institutes of Health (NIH) have created a draft policy that would extend similar reporting requirements to all clinical trials funded by NIH.
About the NPRM: Who, what, and when
The NPRM details new procedures for meeting the requirements established by the Food and Drug Administration Amendments Act of 2007 (FDAAA) to improve public access to clinical trial information.
The proposed rule specifies how information about a clinical trial would need to be submitted to ClinicalTrials.gov. It would not affect requirements for the design or conduct of clinical trials or for the data that must be collected during these trials.
The rule would apply to controlled, interventional studies of drugs, biological products, and devices that are regulated by the FDA. This excludes phase 1 studies of drugs and biological products and feasibility studies of devices.
In general, clinical trials of products regulated by the FDA will meet one or more of the following criteria: include one or more sites in the US; study a product manufactured in the US or its territories and exported for use in a trial outside the US; or be conducted under an FDA investigational new drug application or investigational device exemption.
The NPRM would require the parties responsible for applicable clinical trials—such as the sponsor or a designated principal investigator—to register the trial at ClinicalTrials.gov no later than 21 days after enrolling the first participant.
Registration consists of submitting 4 categories of data elements that are specified in the NPRM: 1) descriptive information, 2) recruitment information, 3) location and contact information, and 4) administrative information.
The parties responsible for the trial would also be required to submit a summary of the study’s results to ClinicalTrials.gov, generally no later than 12 months after trial completion.
However, the NPRM includes procedures for delaying results submission and for requesting extensions to the results submission deadline for good cause.
NIH policy: Extending the NPRM
The proposed NIH policy would make requirements in the NPRM applicable to all NIH-funded awardees and investigators conducting clinical trials, regardless of study phase, type of intervention, or whether they are subject to the rules proposed in the NPRM.
NIH awardees would be expected to ensure submission to ClinicalTrials.gov of the same type of registration and results information, and in the same timeframes, as responsible parties whose trials are subject to the FDAAA and the regulations proposed in the NPRM.
For clinical trials subject to only the proposed NIH policy (not the FDAAA), the NIH would post submitted information, in general, no later than 30 days after it is submitted.
An NIH-funded clinical trial that is also subject to FDAAA would need to have only one entry in ClinicalTrials.gov containing its registration and results information.
Open for comment
The public may comment on any aspect of the NPRM or the proposed NIH policy.
Written comments on the NPRM should be submitted to docket number NIH-2011-0003 at www.regulations.gov. Commenters should indicate the specific section of the NPRM to which each comment refers.
Written comments on the proposed NIH policy should be submitted to the Office of Clinical Research and Bioethics Policy, Office of Science Policy, NIH, via email at clinicaltrials.disseminationpolicy@mail.nih.gov, mail at 6705 Rockledge Drive, Suite 750, Bethesda, MD 20892, or fax at 301-496-9839. ![]()
Studies reveal secrets of malaria transmission
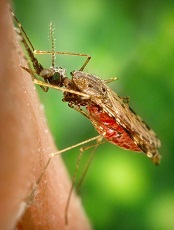
Credit: James Gathany
Two studies comparing mosquito genomes have begun to provide answers to a century-old mystery: why only some Anopheles mosquitoes transmit human malaria.
There are more than 400 species of Anopheles mosquitoes, but only about 60 transmit parasites that cause malaria in humans.
Variation in the ability to transmit malaria, or “vectorial capacity,” is determined by many factors, including feeding and breeding preferences, as well as immune responses to infections.
Much of our understanding of such processes is derived from the sequencing of the Anopheles gambiae genome in 2002, which facilitated large-scale functional studies that have offered insights into how this mosquito became highly specialized to live among and feed upon humans.
The lack of such genomic resources for other Anopheles species limited comparisons to small-scale studies of individual genes with no genome-wide data to investigate key attributes that impact the mosquitoes’ ability to transmit parasites.
In an attempt to change that, Daniel Neafsey, PhD, of the Broad Institute in Cambridge, Massachusetts, and his colleagues sequenced the genomes of 16 anopheline mosquito species.
A second team of researchers—Michael Fontaine, PhD, of the University of Notre Dame in Indiana, and his colleagues—leveraged the 16 reference sequences to uncover additional information.
Both groups described their work in Science Express.
The researchers performed comparative genomics analyses among the Anopheles species and Drosophila (one of the most closely related genera for which equivalent genomic resources exist). This revealed significant genetic differences that make certain Anopheles species particularly adept at inflicting life-threatening infections.
Anopheles species had high rates of gene gain and loss, about 5 times higher than in Drosophila. Some genes, such as those involved in reproduction or those that encode proteins secreted into the mosquito saliva, have very high rates of sequence evolution and are only found in subsets of the most closely related species.
“These dynamic changes may offer clues to understanding the diversification of Anopheles mosquitoes: why some breed in salty water while others need temporary or permanent pools of fresh water, or why some are attracted to livestock while others will only feed on humans,” said Nora Besansky, PhD, a professor at the University of Notre Dame and senior author of both studies.
The genome sequences also provided conclusive evidence of the true relations among several species that are very closely related to Anopheles gambiae but show different traits that affect their vectorial capacity.
“The question of the true species phylogeny has been a highly contentious issue in the field,” Dr Besansky said. “Our results show that the most efficient vectors are not necessarily the most closely related species, and that traits enhancing vectorial capacity may be gained by gene flow between species.”
The researchers found that gene flow is much more extensive in anophelines than in Drosophila, largely because of a process called introgression, whereby a gene from one species enters the gene pool of another. The process allows for a much more rapid transfer of genes than would arise simply by waiting for new mutations to crop up.
The high degree of anopheline gene flow provides a source of genetic variation on which natural selection can act—paving the way for traits that make mosquitoes highly effective vectors for malaria (like insecticide resistance or tolerance of more malaria parasites) to be fixed in certain anophelines. ![]()

Credit: James Gathany
Two studies comparing mosquito genomes have begun to provide answers to a century-old mystery: why only some Anopheles mosquitoes transmit human malaria.
There are more than 400 species of Anopheles mosquitoes, but only about 60 transmit parasites that cause malaria in humans.
Variation in the ability to transmit malaria, or “vectorial capacity,” is determined by many factors, including feeding and breeding preferences, as well as immune responses to infections.
Much of our understanding of such processes is derived from the sequencing of the Anopheles gambiae genome in 2002, which facilitated large-scale functional studies that have offered insights into how this mosquito became highly specialized to live among and feed upon humans.
The lack of such genomic resources for other Anopheles species limited comparisons to small-scale studies of individual genes with no genome-wide data to investigate key attributes that impact the mosquitoes’ ability to transmit parasites.
In an attempt to change that, Daniel Neafsey, PhD, of the Broad Institute in Cambridge, Massachusetts, and his colleagues sequenced the genomes of 16 anopheline mosquito species.
A second team of researchers—Michael Fontaine, PhD, of the University of Notre Dame in Indiana, and his colleagues—leveraged the 16 reference sequences to uncover additional information.
Both groups described their work in Science Express.
The researchers performed comparative genomics analyses among the Anopheles species and Drosophila (one of the most closely related genera for which equivalent genomic resources exist). This revealed significant genetic differences that make certain Anopheles species particularly adept at inflicting life-threatening infections.
Anopheles species had high rates of gene gain and loss, about 5 times higher than in Drosophila. Some genes, such as those involved in reproduction or those that encode proteins secreted into the mosquito saliva, have very high rates of sequence evolution and are only found in subsets of the most closely related species.
“These dynamic changes may offer clues to understanding the diversification of Anopheles mosquitoes: why some breed in salty water while others need temporary or permanent pools of fresh water, or why some are attracted to livestock while others will only feed on humans,” said Nora Besansky, PhD, a professor at the University of Notre Dame and senior author of both studies.
The genome sequences also provided conclusive evidence of the true relations among several species that are very closely related to Anopheles gambiae but show different traits that affect their vectorial capacity.
“The question of the true species phylogeny has been a highly contentious issue in the field,” Dr Besansky said. “Our results show that the most efficient vectors are not necessarily the most closely related species, and that traits enhancing vectorial capacity may be gained by gene flow between species.”
The researchers found that gene flow is much more extensive in anophelines than in Drosophila, largely because of a process called introgression, whereby a gene from one species enters the gene pool of another. The process allows for a much more rapid transfer of genes than would arise simply by waiting for new mutations to crop up.
The high degree of anopheline gene flow provides a source of genetic variation on which natural selection can act—paving the way for traits that make mosquitoes highly effective vectors for malaria (like insecticide resistance or tolerance of more malaria parasites) to be fixed in certain anophelines. ![]()

Credit: James Gathany
Two studies comparing mosquito genomes have begun to provide answers to a century-old mystery: why only some Anopheles mosquitoes transmit human malaria.
There are more than 400 species of Anopheles mosquitoes, but only about 60 transmit parasites that cause malaria in humans.
Variation in the ability to transmit malaria, or “vectorial capacity,” is determined by many factors, including feeding and breeding preferences, as well as immune responses to infections.
Much of our understanding of such processes is derived from the sequencing of the Anopheles gambiae genome in 2002, which facilitated large-scale functional studies that have offered insights into how this mosquito became highly specialized to live among and feed upon humans.
The lack of such genomic resources for other Anopheles species limited comparisons to small-scale studies of individual genes with no genome-wide data to investigate key attributes that impact the mosquitoes’ ability to transmit parasites.
In an attempt to change that, Daniel Neafsey, PhD, of the Broad Institute in Cambridge, Massachusetts, and his colleagues sequenced the genomes of 16 anopheline mosquito species.
A second team of researchers—Michael Fontaine, PhD, of the University of Notre Dame in Indiana, and his colleagues—leveraged the 16 reference sequences to uncover additional information.
Both groups described their work in Science Express.
The researchers performed comparative genomics analyses among the Anopheles species and Drosophila (one of the most closely related genera for which equivalent genomic resources exist). This revealed significant genetic differences that make certain Anopheles species particularly adept at inflicting life-threatening infections.
Anopheles species had high rates of gene gain and loss, about 5 times higher than in Drosophila. Some genes, such as those involved in reproduction or those that encode proteins secreted into the mosquito saliva, have very high rates of sequence evolution and are only found in subsets of the most closely related species.
“These dynamic changes may offer clues to understanding the diversification of Anopheles mosquitoes: why some breed in salty water while others need temporary or permanent pools of fresh water, or why some are attracted to livestock while others will only feed on humans,” said Nora Besansky, PhD, a professor at the University of Notre Dame and senior author of both studies.
The genome sequences also provided conclusive evidence of the true relations among several species that are very closely related to Anopheles gambiae but show different traits that affect their vectorial capacity.
“The question of the true species phylogeny has been a highly contentious issue in the field,” Dr Besansky said. “Our results show that the most efficient vectors are not necessarily the most closely related species, and that traits enhancing vectorial capacity may be gained by gene flow between species.”
The researchers found that gene flow is much more extensive in anophelines than in Drosophila, largely because of a process called introgression, whereby a gene from one species enters the gene pool of another. The process allows for a much more rapid transfer of genes than would arise simply by waiting for new mutations to crop up.
The high degree of anopheline gene flow provides a source of genetic variation on which natural selection can act—paving the way for traits that make mosquitoes highly effective vectors for malaria (like insecticide resistance or tolerance of more malaria parasites) to be fixed in certain anophelines. ![]()
FDA influence on design of pivotal drug studies

Credit: FDA
New research suggests that 20% of recent drug approvals occurred without pharmaceutical companies and the US Food and Drug Administration (FDA) meeting to discuss pivotal studies.
When these meetings did occur, companies did not comply with a quarter of FDA recommendations regarding study design or primary outcome.
Steven Woloshin, MD, of the Dartmouth Institute for Health Policy and Clinical Practice in Lebanon, New Hampshire, and his colleagues reported these findings in JAMA.
The researchers noted that federal regulations encourage but do not require meetings between pharmaceutical companies and the FDA during the design phase of pivotal studies assessing drug efficacy and safety for the proposed indication.
These meetings often generate FDA recommendations for improving research, but companies are not bound to follow them.
To evaluate this process, Dr Woloshin and his colleagues reviewed and analyzed approximately 200 FDA documents (memos, meeting minutes, filing checklists, and medical, statistical, and summary reviews) for 35 new drugs approved between February 1, 2011, and February 29, 2012.
The researchers identified all FDA comments and analyzed recommendations about pivotal study design or primary outcomes and characterized the effect of recommendations on study quality.
Of 35 new drug approvals, companies met with the FDA to discuss pivotal studies for 28 (80%). The FDA made 53 recommendations about design (eg, controls, doses, study length) or primary outcome for 21 approvals.
Fifty-one recommendations were judged as increasing study quality (eg, adding controls, blinding, or specific measures and frequency for toxicity assessments, lengthening studies to assess outcome durability) and 2 as having an uncertain effect.
Companies complied with 40 of the 53 recommendations (75%). An example of non-compliance is the FDA recommending randomized trials of brentuximab and crizotinib, with the companies conducting uncontrolled studies.
Other cases included primary outcome choice (eg, progression-free survival instead of overall survival) and drug (active comparator) doses tested.
Companies can also request FDA review of pivotal trial protocols. If the FDA endorses the protocol, it agrees not to object to any study design issues when reviewing the drug for approval.
Companies requested protocol review for 21 of the 35 new drug approvals, and the FDA endorsed the protocol for 12.
The researchers said instituting mandatory FDA review of pivotal trial protocols with the power to issue binding recommendations could be an effective way to optimize study quality.
An FDA-commissioned report suggested that stronger early FDA involvement could prevent deficiencies that delay the approval of effective drugs and more clearly identify drugs that are ineffective or harmful. ![]()

Credit: FDA
New research suggests that 20% of recent drug approvals occurred without pharmaceutical companies and the US Food and Drug Administration (FDA) meeting to discuss pivotal studies.
When these meetings did occur, companies did not comply with a quarter of FDA recommendations regarding study design or primary outcome.
Steven Woloshin, MD, of the Dartmouth Institute for Health Policy and Clinical Practice in Lebanon, New Hampshire, and his colleagues reported these findings in JAMA.
The researchers noted that federal regulations encourage but do not require meetings between pharmaceutical companies and the FDA during the design phase of pivotal studies assessing drug efficacy and safety for the proposed indication.
These meetings often generate FDA recommendations for improving research, but companies are not bound to follow them.
To evaluate this process, Dr Woloshin and his colleagues reviewed and analyzed approximately 200 FDA documents (memos, meeting minutes, filing checklists, and medical, statistical, and summary reviews) for 35 new drugs approved between February 1, 2011, and February 29, 2012.
The researchers identified all FDA comments and analyzed recommendations about pivotal study design or primary outcomes and characterized the effect of recommendations on study quality.
Of 35 new drug approvals, companies met with the FDA to discuss pivotal studies for 28 (80%). The FDA made 53 recommendations about design (eg, controls, doses, study length) or primary outcome for 21 approvals.
Fifty-one recommendations were judged as increasing study quality (eg, adding controls, blinding, or specific measures and frequency for toxicity assessments, lengthening studies to assess outcome durability) and 2 as having an uncertain effect.
Companies complied with 40 of the 53 recommendations (75%). An example of non-compliance is the FDA recommending randomized trials of brentuximab and crizotinib, with the companies conducting uncontrolled studies.
Other cases included primary outcome choice (eg, progression-free survival instead of overall survival) and drug (active comparator) doses tested.
Companies can also request FDA review of pivotal trial protocols. If the FDA endorses the protocol, it agrees not to object to any study design issues when reviewing the drug for approval.
Companies requested protocol review for 21 of the 35 new drug approvals, and the FDA endorsed the protocol for 12.
The researchers said instituting mandatory FDA review of pivotal trial protocols with the power to issue binding recommendations could be an effective way to optimize study quality.
An FDA-commissioned report suggested that stronger early FDA involvement could prevent deficiencies that delay the approval of effective drugs and more clearly identify drugs that are ineffective or harmful. ![]()

Credit: FDA
New research suggests that 20% of recent drug approvals occurred without pharmaceutical companies and the US Food and Drug Administration (FDA) meeting to discuss pivotal studies.
When these meetings did occur, companies did not comply with a quarter of FDA recommendations regarding study design or primary outcome.
Steven Woloshin, MD, of the Dartmouth Institute for Health Policy and Clinical Practice in Lebanon, New Hampshire, and his colleagues reported these findings in JAMA.
The researchers noted that federal regulations encourage but do not require meetings between pharmaceutical companies and the FDA during the design phase of pivotal studies assessing drug efficacy and safety for the proposed indication.
These meetings often generate FDA recommendations for improving research, but companies are not bound to follow them.
To evaluate this process, Dr Woloshin and his colleagues reviewed and analyzed approximately 200 FDA documents (memos, meeting minutes, filing checklists, and medical, statistical, and summary reviews) for 35 new drugs approved between February 1, 2011, and February 29, 2012.
The researchers identified all FDA comments and analyzed recommendations about pivotal study design or primary outcomes and characterized the effect of recommendations on study quality.
Of 35 new drug approvals, companies met with the FDA to discuss pivotal studies for 28 (80%). The FDA made 53 recommendations about design (eg, controls, doses, study length) or primary outcome for 21 approvals.
Fifty-one recommendations were judged as increasing study quality (eg, adding controls, blinding, or specific measures and frequency for toxicity assessments, lengthening studies to assess outcome durability) and 2 as having an uncertain effect.
Companies complied with 40 of the 53 recommendations (75%). An example of non-compliance is the FDA recommending randomized trials of brentuximab and crizotinib, with the companies conducting uncontrolled studies.
Other cases included primary outcome choice (eg, progression-free survival instead of overall survival) and drug (active comparator) doses tested.
Companies can also request FDA review of pivotal trial protocols. If the FDA endorses the protocol, it agrees not to object to any study design issues when reviewing the drug for approval.
Companies requested protocol review for 21 of the 35 new drug approvals, and the FDA endorsed the protocol for 12.
The researchers said instituting mandatory FDA review of pivotal trial protocols with the power to issue binding recommendations could be an effective way to optimize study quality.
An FDA-commissioned report suggested that stronger early FDA involvement could prevent deficiencies that delay the approval of effective drugs and more clearly identify drugs that are ineffective or harmful.
Map helps predict new cancer genes

Credit: Mount Sinai Hospital
Researchers say they’ve created the largest-scale map of direct interactions between proteins encoded by the human genome, and this has revealed dozens of genes that may be involved in cancers.
This human interactome map describes about 14,000 direct interactions between proteins.
The map is about 30% larger than previous maps and contains more high-quality interactions than have come from all previous studies combined, according to the researchers.
Frederick Roth, PhD, of the University of Toronto and Mount Sinai Hospital in Ontario, Canada, and his colleagues described their map in Cell.
First, the researchers identified protein interactions via lab experiments. Then, they used computer modelling to zoom in on proteins that connect to one or more other cancer proteins.
“We show, really for the first time, that cancer proteins are more likely to interconnect with one another than they are to connect to randomly chosen non-cancer proteins,” Dr Roth said.
“Once you see that proteins associated to the same disease are more likely to connect to each other, now you can use this network of interactions as a prediction tool to find new cancer proteins and the genes they encode.”
For example, two known cancer genes encoded two proteins that interacted with CTBP2, a protein encoded at a location tied to prostate cancer, which can spread to nearby lymph nodes. These two proteins are implicated in lymphoid tumors, suggesting that CTBP2 plays a role in the development of lymphoid tumors.
Using their predictive method, the researchers found that 60 of their predicted cancer genes fit into a known cancer pathway.
The study also revealed that the network of protein interactions in humans covers a much broader range of genes than some past research has suggested.
Dr Roth said studies often focus on “popular” proteins that have already been linked to disease or are interesting for other reasons, and this has created a bias in our understanding of protein interactions.
“One major conclusion of the paper is that when you look systematically for interactions, you find them everywhere,” he said.
He and his colleagues believe that knowledge of protein interactions is likely to inform worldwide efforts to sequence and interpret cancer genomes.

Credit: Mount Sinai Hospital
Researchers say they’ve created the largest-scale map of direct interactions between proteins encoded by the human genome, and this has revealed dozens of genes that may be involved in cancers.
This human interactome map describes about 14,000 direct interactions between proteins.
The map is about 30% larger than previous maps and contains more high-quality interactions than have come from all previous studies combined, according to the researchers.
Frederick Roth, PhD, of the University of Toronto and Mount Sinai Hospital in Ontario, Canada, and his colleagues described their map in Cell.
First, the researchers identified protein interactions via lab experiments. Then, they used computer modelling to zoom in on proteins that connect to one or more other cancer proteins.
“We show, really for the first time, that cancer proteins are more likely to interconnect with one another than they are to connect to randomly chosen non-cancer proteins,” Dr Roth said.
“Once you see that proteins associated to the same disease are more likely to connect to each other, now you can use this network of interactions as a prediction tool to find new cancer proteins and the genes they encode.”
For example, two known cancer genes encoded two proteins that interacted with CTBP2, a protein encoded at a location tied to prostate cancer, which can spread to nearby lymph nodes. These two proteins are implicated in lymphoid tumors, suggesting that CTBP2 plays a role in the development of lymphoid tumors.
Using their predictive method, the researchers found that 60 of their predicted cancer genes fit into a known cancer pathway.
The study also revealed that the network of protein interactions in humans covers a much broader range of genes than some past research has suggested.
Dr Roth said studies often focus on “popular” proteins that have already been linked to disease or are interesting for other reasons, and this has created a bias in our understanding of protein interactions.
“One major conclusion of the paper is that when you look systematically for interactions, you find them everywhere,” he said.
He and his colleagues believe that knowledge of protein interactions is likely to inform worldwide efforts to sequence and interpret cancer genomes.

Credit: Mount Sinai Hospital
Researchers say they’ve created the largest-scale map of direct interactions between proteins encoded by the human genome, and this has revealed dozens of genes that may be involved in cancers.
This human interactome map describes about 14,000 direct interactions between proteins.
The map is about 30% larger than previous maps and contains more high-quality interactions than have come from all previous studies combined, according to the researchers.
Frederick Roth, PhD, of the University of Toronto and Mount Sinai Hospital in Ontario, Canada, and his colleagues described their map in Cell.
First, the researchers identified protein interactions via lab experiments. Then, they used computer modelling to zoom in on proteins that connect to one or more other cancer proteins.
“We show, really for the first time, that cancer proteins are more likely to interconnect with one another than they are to connect to randomly chosen non-cancer proteins,” Dr Roth said.
“Once you see that proteins associated to the same disease are more likely to connect to each other, now you can use this network of interactions as a prediction tool to find new cancer proteins and the genes they encode.”
For example, two known cancer genes encoded two proteins that interacted with CTBP2, a protein encoded at a location tied to prostate cancer, which can spread to nearby lymph nodes. These two proteins are implicated in lymphoid tumors, suggesting that CTBP2 plays a role in the development of lymphoid tumors.
Using their predictive method, the researchers found that 60 of their predicted cancer genes fit into a known cancer pathway.
The study also revealed that the network of protein interactions in humans covers a much broader range of genes than some past research has suggested.
Dr Roth said studies often focus on “popular” proteins that have already been linked to disease or are interesting for other reasons, and this has created a bias in our understanding of protein interactions.
“One major conclusion of the paper is that when you look systematically for interactions, you find them everywhere,” he said.
He and his colleagues believe that knowledge of protein interactions is likely to inform worldwide efforts to sequence and interpret cancer genomes.