User login
Tool mines scientific literature, generates hypotheses

Credit: NIH
A new tool may help researchers sift through the scientific literature to discover hypothesis-generating information relevant to their own research.
The resource, called the Knowledge Integration Toolkit (KnIT), extracts relevant information from the literature, includes it in a network that can be queried, and
then attempts to use these data to generate reasonable and testable hypotheses that can help direct lab studies.
Researchers tested KnIT in a retrospective case study involving published data on p53 and found the tool could accurately predict the existence of proteins that modify p53.
Details from this study were published in the Association for Computing Machinery’s digital library.
Olivier Lichtarge, MD, PhD, of the Baylor College of Medicine in Houston, Texas, is scheduled to discuss the study on August 27 at the 20th Annual Association for Computing Machinery’s Special Interest Group on Knowledge Discovery and Data Mining Conference in New York, New York.
“On average, a scientist might read between 1 and 5 research papers on a good day,” Dr Lichtarge noted.
“But, to put this in perspective with p53, there are over 70,000 papers published on this protein. Even if a scientist reads 5 papers a day, it could take nearly 38 years to completely understand all of the research already available today on this protein.”
Scientists formulate hypotheses based on what they read and know, but because they cannot read everything, their hypotheses may be biased, according to Dr Lichtarge.
“A computer certainly may not reason as well as a scientist,” he said, “but the little it can, logically and objectively, may contribute greatly when applied to our entire body of knowledge.”
With that in mind, Dr Lichtarge and his colleagues initiated a project to develop a knowledge integration tool that took advantage of existing text mining capabilities, such as those used by IBM’s Watson technology—cognitive technology that processes information more like a human than a computer.
And the team came up with KnIT. In the first test using KnIT, they sought to identify new protein kinases that phosphorylate p53.
There are more than 500 known human kinases and tens of thousands of possible proteins they can target. Thirty-three are currently known to modify p53.
The researchers used KnIT to mine the scientific literature up to 2003, when only half of the 33 phosphorylating protein kinases had been discovered.
Seventy-four kinases were extracted as potential modifiers. Of these, prior to 2003, 10 were known to phosphorylate p53, and 9 were discovered at a later date.
Of the 10 already known, KnIT accounted for them in reasoning as well as ranking the likelihood that the other 64 kinases targeted p53. Of the 9 found nearly a decade later, KnIT accurately predicted 7.
“This study showed that, in a very narrow field of study regarding p53, we can, in fact, suggest new relationships and new functions associated with p53, which can later be directly validated in the laboratory,” Dr Lichtarge said.
“Our long-term hope is to systematically extract knowledge directly from the totality of the public medical literature. For this, we need technological advances to read text, extract facts from every sentence, and to integrate this information into a network that describes the relationship between all of the objects and entities discussed in the literature.”
“This first study is promising, because it suggests a proof of principle for a small step towards this type of knowledge discovery. With more research, we hope to get closer to clinical and therapeutic applications.” ![]()

Credit: NIH
A new tool may help researchers sift through the scientific literature to discover hypothesis-generating information relevant to their own research.
The resource, called the Knowledge Integration Toolkit (KnIT), extracts relevant information from the literature, includes it in a network that can be queried, and
then attempts to use these data to generate reasonable and testable hypotheses that can help direct lab studies.
Researchers tested KnIT in a retrospective case study involving published data on p53 and found the tool could accurately predict the existence of proteins that modify p53.
Details from this study were published in the Association for Computing Machinery’s digital library.
Olivier Lichtarge, MD, PhD, of the Baylor College of Medicine in Houston, Texas, is scheduled to discuss the study on August 27 at the 20th Annual Association for Computing Machinery’s Special Interest Group on Knowledge Discovery and Data Mining Conference in New York, New York.
“On average, a scientist might read between 1 and 5 research papers on a good day,” Dr Lichtarge noted.
“But, to put this in perspective with p53, there are over 70,000 papers published on this protein. Even if a scientist reads 5 papers a day, it could take nearly 38 years to completely understand all of the research already available today on this protein.”
Scientists formulate hypotheses based on what they read and know, but because they cannot read everything, their hypotheses may be biased, according to Dr Lichtarge.
“A computer certainly may not reason as well as a scientist,” he said, “but the little it can, logically and objectively, may contribute greatly when applied to our entire body of knowledge.”
With that in mind, Dr Lichtarge and his colleagues initiated a project to develop a knowledge integration tool that took advantage of existing text mining capabilities, such as those used by IBM’s Watson technology—cognitive technology that processes information more like a human than a computer.
And the team came up with KnIT. In the first test using KnIT, they sought to identify new protein kinases that phosphorylate p53.
There are more than 500 known human kinases and tens of thousands of possible proteins they can target. Thirty-three are currently known to modify p53.
The researchers used KnIT to mine the scientific literature up to 2003, when only half of the 33 phosphorylating protein kinases had been discovered.
Seventy-four kinases were extracted as potential modifiers. Of these, prior to 2003, 10 were known to phosphorylate p53, and 9 were discovered at a later date.
Of the 10 already known, KnIT accounted for them in reasoning as well as ranking the likelihood that the other 64 kinases targeted p53. Of the 9 found nearly a decade later, KnIT accurately predicted 7.
“This study showed that, in a very narrow field of study regarding p53, we can, in fact, suggest new relationships and new functions associated with p53, which can later be directly validated in the laboratory,” Dr Lichtarge said.
“Our long-term hope is to systematically extract knowledge directly from the totality of the public medical literature. For this, we need technological advances to read text, extract facts from every sentence, and to integrate this information into a network that describes the relationship between all of the objects and entities discussed in the literature.”
“This first study is promising, because it suggests a proof of principle for a small step towards this type of knowledge discovery. With more research, we hope to get closer to clinical and therapeutic applications.” ![]()

Credit: NIH
A new tool may help researchers sift through the scientific literature to discover hypothesis-generating information relevant to their own research.
The resource, called the Knowledge Integration Toolkit (KnIT), extracts relevant information from the literature, includes it in a network that can be queried, and
then attempts to use these data to generate reasonable and testable hypotheses that can help direct lab studies.
Researchers tested KnIT in a retrospective case study involving published data on p53 and found the tool could accurately predict the existence of proteins that modify p53.
Details from this study were published in the Association for Computing Machinery’s digital library.
Olivier Lichtarge, MD, PhD, of the Baylor College of Medicine in Houston, Texas, is scheduled to discuss the study on August 27 at the 20th Annual Association for Computing Machinery’s Special Interest Group on Knowledge Discovery and Data Mining Conference in New York, New York.
“On average, a scientist might read between 1 and 5 research papers on a good day,” Dr Lichtarge noted.
“But, to put this in perspective with p53, there are over 70,000 papers published on this protein. Even if a scientist reads 5 papers a day, it could take nearly 38 years to completely understand all of the research already available today on this protein.”
Scientists formulate hypotheses based on what they read and know, but because they cannot read everything, their hypotheses may be biased, according to Dr Lichtarge.
“A computer certainly may not reason as well as a scientist,” he said, “but the little it can, logically and objectively, may contribute greatly when applied to our entire body of knowledge.”
With that in mind, Dr Lichtarge and his colleagues initiated a project to develop a knowledge integration tool that took advantage of existing text mining capabilities, such as those used by IBM’s Watson technology—cognitive technology that processes information more like a human than a computer.
And the team came up with KnIT. In the first test using KnIT, they sought to identify new protein kinases that phosphorylate p53.
There are more than 500 known human kinases and tens of thousands of possible proteins they can target. Thirty-three are currently known to modify p53.
The researchers used KnIT to mine the scientific literature up to 2003, when only half of the 33 phosphorylating protein kinases had been discovered.
Seventy-four kinases were extracted as potential modifiers. Of these, prior to 2003, 10 were known to phosphorylate p53, and 9 were discovered at a later date.
Of the 10 already known, KnIT accounted for them in reasoning as well as ranking the likelihood that the other 64 kinases targeted p53. Of the 9 found nearly a decade later, KnIT accurately predicted 7.
“This study showed that, in a very narrow field of study regarding p53, we can, in fact, suggest new relationships and new functions associated with p53, which can later be directly validated in the laboratory,” Dr Lichtarge said.
“Our long-term hope is to systematically extract knowledge directly from the totality of the public medical literature. For this, we need technological advances to read text, extract facts from every sentence, and to integrate this information into a network that describes the relationship between all of the objects and entities discussed in the literature.”
“This first study is promising, because it suggests a proof of principle for a small step towards this type of knowledge discovery. With more research, we hope to get closer to clinical and therapeutic applications.” ![]()
A new system for malaria diagnosis
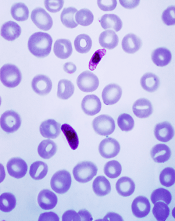
Plasmodium falciparum
Credit: CDC/Mae Melvin
A semi-automated system may allow healthcare professionals to diagnose malaria infection with more than 90% accuracy.
With this system, a computer algorithm analyzes red blood cells from a digitized slide, ranks them according to the likelihood of infection, and presents the user with more than 100 images from which to make a diagnosis.
Johan Lundin, MD, PhD, of the Institute for Molecular Medicine Finland, and his colleagues described the system in PLOS ONE.
The researchers noted that high-quality microscopy is still the most accurate method for detecting malaria infection. However, as microscopy can be time-consuming, the team wanted to streamline the process by developing a semi-automated system.
“We are not suggesting that the whole malaria diagnostic process could or should be automated,” Dr Lundin said. “Rather, our aim is to develop methods that are significantly less labor-intensive than the traditional ones and have a potential to considerably increase the throughput in malaria diagnostics.”
The group’s method is based on computer vision algorithms similar to those used in facial recognition software. First, a thin layer of blood smeared on a microscope slide is digitized. Then, a computer algorithm analyzes more than 50,000 red blood cells per sample and ranks them according to the probability of malaria infection.
Next, the program creates a panel containing images of more than a hundred cells that are the most likely to be infected and presents that panel to the user. The final diagnosis is made by a healthcare professional based on the images.
To test this system, Dr Lundin and his colleagues used a set of samples from 19 patients already diagnosed with malaria and 12 control subjects. From each sample, the researchers created a digitized slide, and the system generated 128 images of the most probable parasite candidate regions.
Two expert microscopists viewed the images on a tablet computer to determine whether a subject was infected with Plasmodium falciparum.
The diagnostic sensitivity was 90% with one viewer and 95% for the other. The specificity was 100% for both viewers.
Based on these results, the researchers said this system has the potential to increase the throughput in malaria diagnostics. However, it does require some tweaking, and the team would like to expand its capabilities.
“The equipment needed for digitization of the samples is a challenge in developed countries,” said study author Nina Linder, MD, PhD, also of the Institute for Molecular Medicine Finland. “In the next phase of our project, we will test the system in combination with inexpensive mobile microscopy devices that our group has also developed.”
“There is also a strong need for fast and accurate methods for measuring the malaria parasite load in a sample,” she added. “Various malaria drug screening programs are underway, and the parasite load in a large number of samples needs to be quantified for determining the efficacy of potential drugs. We are further developing the computer algorithms used in this study to meet this need as well.”
Lastly, the researchers said this system could be applied in various other fields of medicine. In addition to other infectious diseases such as tuberculosis, the group is planning to test the system’s utility in cancer diagnosis. ![]()

Plasmodium falciparum
Credit: CDC/Mae Melvin
A semi-automated system may allow healthcare professionals to diagnose malaria infection with more than 90% accuracy.
With this system, a computer algorithm analyzes red blood cells from a digitized slide, ranks them according to the likelihood of infection, and presents the user with more than 100 images from which to make a diagnosis.
Johan Lundin, MD, PhD, of the Institute for Molecular Medicine Finland, and his colleagues described the system in PLOS ONE.
The researchers noted that high-quality microscopy is still the most accurate method for detecting malaria infection. However, as microscopy can be time-consuming, the team wanted to streamline the process by developing a semi-automated system.
“We are not suggesting that the whole malaria diagnostic process could or should be automated,” Dr Lundin said. “Rather, our aim is to develop methods that are significantly less labor-intensive than the traditional ones and have a potential to considerably increase the throughput in malaria diagnostics.”
The group’s method is based on computer vision algorithms similar to those used in facial recognition software. First, a thin layer of blood smeared on a microscope slide is digitized. Then, a computer algorithm analyzes more than 50,000 red blood cells per sample and ranks them according to the probability of malaria infection.
Next, the program creates a panel containing images of more than a hundred cells that are the most likely to be infected and presents that panel to the user. The final diagnosis is made by a healthcare professional based on the images.
To test this system, Dr Lundin and his colleagues used a set of samples from 19 patients already diagnosed with malaria and 12 control subjects. From each sample, the researchers created a digitized slide, and the system generated 128 images of the most probable parasite candidate regions.
Two expert microscopists viewed the images on a tablet computer to determine whether a subject was infected with Plasmodium falciparum.
The diagnostic sensitivity was 90% with one viewer and 95% for the other. The specificity was 100% for both viewers.
Based on these results, the researchers said this system has the potential to increase the throughput in malaria diagnostics. However, it does require some tweaking, and the team would like to expand its capabilities.
“The equipment needed for digitization of the samples is a challenge in developed countries,” said study author Nina Linder, MD, PhD, also of the Institute for Molecular Medicine Finland. “In the next phase of our project, we will test the system in combination with inexpensive mobile microscopy devices that our group has also developed.”
“There is also a strong need for fast and accurate methods for measuring the malaria parasite load in a sample,” she added. “Various malaria drug screening programs are underway, and the parasite load in a large number of samples needs to be quantified for determining the efficacy of potential drugs. We are further developing the computer algorithms used in this study to meet this need as well.”
Lastly, the researchers said this system could be applied in various other fields of medicine. In addition to other infectious diseases such as tuberculosis, the group is planning to test the system’s utility in cancer diagnosis. ![]()

Plasmodium falciparum
Credit: CDC/Mae Melvin
A semi-automated system may allow healthcare professionals to diagnose malaria infection with more than 90% accuracy.
With this system, a computer algorithm analyzes red blood cells from a digitized slide, ranks them according to the likelihood of infection, and presents the user with more than 100 images from which to make a diagnosis.
Johan Lundin, MD, PhD, of the Institute for Molecular Medicine Finland, and his colleagues described the system in PLOS ONE.
The researchers noted that high-quality microscopy is still the most accurate method for detecting malaria infection. However, as microscopy can be time-consuming, the team wanted to streamline the process by developing a semi-automated system.
“We are not suggesting that the whole malaria diagnostic process could or should be automated,” Dr Lundin said. “Rather, our aim is to develop methods that are significantly less labor-intensive than the traditional ones and have a potential to considerably increase the throughput in malaria diagnostics.”
The group’s method is based on computer vision algorithms similar to those used in facial recognition software. First, a thin layer of blood smeared on a microscope slide is digitized. Then, a computer algorithm analyzes more than 50,000 red blood cells per sample and ranks them according to the probability of malaria infection.
Next, the program creates a panel containing images of more than a hundred cells that are the most likely to be infected and presents that panel to the user. The final diagnosis is made by a healthcare professional based on the images.
To test this system, Dr Lundin and his colleagues used a set of samples from 19 patients already diagnosed with malaria and 12 control subjects. From each sample, the researchers created a digitized slide, and the system generated 128 images of the most probable parasite candidate regions.
Two expert microscopists viewed the images on a tablet computer to determine whether a subject was infected with Plasmodium falciparum.
The diagnostic sensitivity was 90% with one viewer and 95% for the other. The specificity was 100% for both viewers.
Based on these results, the researchers said this system has the potential to increase the throughput in malaria diagnostics. However, it does require some tweaking, and the team would like to expand its capabilities.
“The equipment needed for digitization of the samples is a challenge in developed countries,” said study author Nina Linder, MD, PhD, also of the Institute for Molecular Medicine Finland. “In the next phase of our project, we will test the system in combination with inexpensive mobile microscopy devices that our group has also developed.”
“There is also a strong need for fast and accurate methods for measuring the malaria parasite load in a sample,” she added. “Various malaria drug screening programs are underway, and the parasite load in a large number of samples needs to be quantified for determining the efficacy of potential drugs. We are further developing the computer algorithms used in this study to meet this need as well.”
Lastly, the researchers said this system could be applied in various other fields of medicine. In addition to other infectious diseases such as tuberculosis, the group is planning to test the system’s utility in cancer diagnosis. ![]()
New technology enables malaria discovery
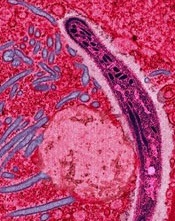
Credit: Ute Frevert
and Margaret Shear
Researchers say they’ve developed a new computational method to predict the function of disease-causing genes and proteins, and this has enabled an important discovery.
The team used the method to study EXP1, a protein known to be essential to the malaria parasite Plasmodium falciparum, although its exact mechanism has been unclear.
The research revealed that EXP1 enables the parasite to detoxify the main metabolic byproducts it creates in red blood cells.
The researchers also found that EXP1 has a direct role in drug action and susceptibility to artesunate, an important member of the artemisinin drug family.
“Through this multiyear collaborative effort, we now have an improved understanding of the protective molecular mechanisms of the malaria parasite and its drug susceptibility to artesunate,” said study author Olivier Lichtarge, MD, PhD, of the Baylor College of Medicine in Houston, Texas.
“As we are witnessing a rise of resistance to artemisinins, these results may help [us in] finding new pathways to successor drugs.”
Dr Lichtarge and his colleagues described this research in Cell.
The team devised a computational method that allows biological information to flow from gene to gene across the supergenomic network.
“The network connects millions of genes from hundreds of species based on their interactions within the organism or based on their ancestral relations between different species,” said study author Andreas Martin Lisewski, PhD, also of the Baylor College of Medicine.
“Normally, computing the flow of functional information would be costly and slow, but we developed a compression method that reduces this gigantic network into one that is much smaller and now computationally tractable. The surprise is that these biological networks are compressible, much like digital data in today’s computers.”
To test their method, the researchers looked at functional predictions of P falciparum. It’s been more than 10 years since this parasite’s genome was fully sequenced, but little is known about the function for most of its genes.
“To better understand this disease, we need to identify more functions of the parasite’s genes,” Dr Lisewski said. “This understanding may eventually help us to stem the rise of drug-resistant malaria, such as the emerging resistance to artemisinins.”
With that in mind, the researchers used their computational method to study EXP1. And they discovered the protein is a membrane glutathione S-transferase that degrades cytotoxic hematin but is inhibited by artesunate. ![]()

Credit: Ute Frevert
and Margaret Shear
Researchers say they’ve developed a new computational method to predict the function of disease-causing genes and proteins, and this has enabled an important discovery.
The team used the method to study EXP1, a protein known to be essential to the malaria parasite Plasmodium falciparum, although its exact mechanism has been unclear.
The research revealed that EXP1 enables the parasite to detoxify the main metabolic byproducts it creates in red blood cells.
The researchers also found that EXP1 has a direct role in drug action and susceptibility to artesunate, an important member of the artemisinin drug family.
“Through this multiyear collaborative effort, we now have an improved understanding of the protective molecular mechanisms of the malaria parasite and its drug susceptibility to artesunate,” said study author Olivier Lichtarge, MD, PhD, of the Baylor College of Medicine in Houston, Texas.
“As we are witnessing a rise of resistance to artemisinins, these results may help [us in] finding new pathways to successor drugs.”
Dr Lichtarge and his colleagues described this research in Cell.
The team devised a computational method that allows biological information to flow from gene to gene across the supergenomic network.
“The network connects millions of genes from hundreds of species based on their interactions within the organism or based on their ancestral relations between different species,” said study author Andreas Martin Lisewski, PhD, also of the Baylor College of Medicine.
“Normally, computing the flow of functional information would be costly and slow, but we developed a compression method that reduces this gigantic network into one that is much smaller and now computationally tractable. The surprise is that these biological networks are compressible, much like digital data in today’s computers.”
To test their method, the researchers looked at functional predictions of P falciparum. It’s been more than 10 years since this parasite’s genome was fully sequenced, but little is known about the function for most of its genes.
“To better understand this disease, we need to identify more functions of the parasite’s genes,” Dr Lisewski said. “This understanding may eventually help us to stem the rise of drug-resistant malaria, such as the emerging resistance to artemisinins.”
With that in mind, the researchers used their computational method to study EXP1. And they discovered the protein is a membrane glutathione S-transferase that degrades cytotoxic hematin but is inhibited by artesunate. ![]()

Credit: Ute Frevert
and Margaret Shear
Researchers say they’ve developed a new computational method to predict the function of disease-causing genes and proteins, and this has enabled an important discovery.
The team used the method to study EXP1, a protein known to be essential to the malaria parasite Plasmodium falciparum, although its exact mechanism has been unclear.
The research revealed that EXP1 enables the parasite to detoxify the main metabolic byproducts it creates in red blood cells.
The researchers also found that EXP1 has a direct role in drug action and susceptibility to artesunate, an important member of the artemisinin drug family.
“Through this multiyear collaborative effort, we now have an improved understanding of the protective molecular mechanisms of the malaria parasite and its drug susceptibility to artesunate,” said study author Olivier Lichtarge, MD, PhD, of the Baylor College of Medicine in Houston, Texas.
“As we are witnessing a rise of resistance to artemisinins, these results may help [us in] finding new pathways to successor drugs.”
Dr Lichtarge and his colleagues described this research in Cell.
The team devised a computational method that allows biological information to flow from gene to gene across the supergenomic network.
“The network connects millions of genes from hundreds of species based on their interactions within the organism or based on their ancestral relations between different species,” said study author Andreas Martin Lisewski, PhD, also of the Baylor College of Medicine.
“Normally, computing the flow of functional information would be costly and slow, but we developed a compression method that reduces this gigantic network into one that is much smaller and now computationally tractable. The surprise is that these biological networks are compressible, much like digital data in today’s computers.”
To test their method, the researchers looked at functional predictions of P falciparum. It’s been more than 10 years since this parasite’s genome was fully sequenced, but little is known about the function for most of its genes.
“To better understand this disease, we need to identify more functions of the parasite’s genes,” Dr Lisewski said. “This understanding may eventually help us to stem the rise of drug-resistant malaria, such as the emerging resistance to artemisinins.”
With that in mind, the researchers used their computational method to study EXP1. And they discovered the protein is a membrane glutathione S-transferase that degrades cytotoxic hematin but is inhibited by artesunate. ![]()
Path to drug development often not straightforward, study shows

Credit: Rhoda Baer
An analysis of university discoveries licensed to biotechnology firms has revealed early bottlenecks in the drug development process.
Typically, universities do most of the basic research and then license a discovery to a small biotech firm that advances the research. The small firm will then sublicense the discovery to a large firm that can run clinical trials.
But an analysis published in Science Translational Medicine suggests the process rarely follows this straightforward path.
Instead, it often zigzags across biotech firms and between research areas before a drug is finally commercialized.
“The timeline for commercialization is much longer than most people think,” said study author Jerry Thursby, PhD, of the Georgia Institute of Technology in Atlanta.
To study the path of drug development, Dr Thursby and his colleagues built a database of 835 patents in 342 university licenses with biotech firms.
They then traced the path of patents to document whether the inventions were sublicensed to another firm for testing in a new disease category or whether the sublicense was to a large firm for clinical trials or marketing.
In all, 27% of inventions appeared in a second license (sublicense). The average time between invention and first license was 5.5 years, and the average time between first and second license was 3.5 years.
This time span is substantial, the researchers said, given that the average time from discovery to drug approval in the US is 13 years.
The team also found that sublicensing often resets the development timeline because a drug must be tested for an entirely new indication or several new indications.
The disease categories in the licenses analyzed spanned 20 distinct indications, and individual licenses included up to 5 indications. But the categories were very broad, such as “cancer” or “infectious diseases.”
Nevertheless, the researchers saw substantial changes in disease indications from the first license to the second. Only 19% of the inventions remained completely unchanged between the first and second license.
For 44% of inventions, none of the first-license indications remained in the second license. Twenty-eight percent of inventions had indications added between the first and second license, and 9% had indications subtracted.
The researchers said these results suggest a need for policies and initiatives that enhance early translation by more efficiently driving more inventions into multiple disease pipelines.
One option might be the formation of an open-source translational research database that complements clinicaltrials.gov, where patents and licenses for biomedical research thought to be destined for eventual therapeutic use would be logged and shared. ![]()

Credit: Rhoda Baer
An analysis of university discoveries licensed to biotechnology firms has revealed early bottlenecks in the drug development process.
Typically, universities do most of the basic research and then license a discovery to a small biotech firm that advances the research. The small firm will then sublicense the discovery to a large firm that can run clinical trials.
But an analysis published in Science Translational Medicine suggests the process rarely follows this straightforward path.
Instead, it often zigzags across biotech firms and between research areas before a drug is finally commercialized.
“The timeline for commercialization is much longer than most people think,” said study author Jerry Thursby, PhD, of the Georgia Institute of Technology in Atlanta.
To study the path of drug development, Dr Thursby and his colleagues built a database of 835 patents in 342 university licenses with biotech firms.
They then traced the path of patents to document whether the inventions were sublicensed to another firm for testing in a new disease category or whether the sublicense was to a large firm for clinical trials or marketing.
In all, 27% of inventions appeared in a second license (sublicense). The average time between invention and first license was 5.5 years, and the average time between first and second license was 3.5 years.
This time span is substantial, the researchers said, given that the average time from discovery to drug approval in the US is 13 years.
The team also found that sublicensing often resets the development timeline because a drug must be tested for an entirely new indication or several new indications.
The disease categories in the licenses analyzed spanned 20 distinct indications, and individual licenses included up to 5 indications. But the categories were very broad, such as “cancer” or “infectious diseases.”
Nevertheless, the researchers saw substantial changes in disease indications from the first license to the second. Only 19% of the inventions remained completely unchanged between the first and second license.
For 44% of inventions, none of the first-license indications remained in the second license. Twenty-eight percent of inventions had indications added between the first and second license, and 9% had indications subtracted.
The researchers said these results suggest a need for policies and initiatives that enhance early translation by more efficiently driving more inventions into multiple disease pipelines.
One option might be the formation of an open-source translational research database that complements clinicaltrials.gov, where patents and licenses for biomedical research thought to be destined for eventual therapeutic use would be logged and shared. ![]()

Credit: Rhoda Baer
An analysis of university discoveries licensed to biotechnology firms has revealed early bottlenecks in the drug development process.
Typically, universities do most of the basic research and then license a discovery to a small biotech firm that advances the research. The small firm will then sublicense the discovery to a large firm that can run clinical trials.
But an analysis published in Science Translational Medicine suggests the process rarely follows this straightforward path.
Instead, it often zigzags across biotech firms and between research areas before a drug is finally commercialized.
“The timeline for commercialization is much longer than most people think,” said study author Jerry Thursby, PhD, of the Georgia Institute of Technology in Atlanta.
To study the path of drug development, Dr Thursby and his colleagues built a database of 835 patents in 342 university licenses with biotech firms.
They then traced the path of patents to document whether the inventions were sublicensed to another firm for testing in a new disease category or whether the sublicense was to a large firm for clinical trials or marketing.
In all, 27% of inventions appeared in a second license (sublicense). The average time between invention and first license was 5.5 years, and the average time between first and second license was 3.5 years.
This time span is substantial, the researchers said, given that the average time from discovery to drug approval in the US is 13 years.
The team also found that sublicensing often resets the development timeline because a drug must be tested for an entirely new indication or several new indications.
The disease categories in the licenses analyzed spanned 20 distinct indications, and individual licenses included up to 5 indications. But the categories were very broad, such as “cancer” or “infectious diseases.”
Nevertheless, the researchers saw substantial changes in disease indications from the first license to the second. Only 19% of the inventions remained completely unchanged between the first and second license.
For 44% of inventions, none of the first-license indications remained in the second license. Twenty-eight percent of inventions had indications added between the first and second license, and 9% had indications subtracted.
The researchers said these results suggest a need for policies and initiatives that enhance early translation by more efficiently driving more inventions into multiple disease pipelines.
One option might be the formation of an open-source translational research database that complements clinicaltrials.gov, where patents and licenses for biomedical research thought to be destined for eventual therapeutic use would be logged and shared. ![]()
New pipette can move single cells

Credit: Rhoda Baer
Researchers say they have developed a pipette that can transfer a single cell at a time.
The device, called the handheld single-cell pipette (hSCP), has 2 plungers. The first is used to wash and capture a single cell, and the second can release the cell in the desired location.
Lidong Qin, PhD, of Houston Methodist Research Institute in Texas, and his colleagues described the hSCP and reported preliminary results with the device in the Journal of the American Chemical Society.
“Studying single cells and their unique functions has become a frontier in current biomedical research,” Dr Qin said. “One of the biggest challenges for single-cell research is picking out only one cell from a collection of millions of cells.”
He noted that current techniques for withdrawing single cells from a tube or Petri dish can be cumbersome, expensive, and time-consuming.
“Some old and clumsy methods are used to capture single cells,” he explained. “Some researchers use their mouths at one end of the pipette, driven by their own mouth force, to try to ensure only a minimum amount of cell suspension collected. The sample is then checked with a microscope to find out the number of cells captured. The opportunity to get only one cell is hit-or-miss and a bit troublesome.”
“One company provides a million-dollar machine that can help biologists transfer single cells to 96-well plates. Each run costs an additional $1000 to purchase the plate. Such technology will not be widely accessible to biologists.”
With that in mind, Dr Qin and his colleagues developed their 2-plunger hSCP. The first plunger withdraws fluid from a suspension of cells.
Fluid travels through canals on either side of a nanoscopic, laser-sculpted “hook” that is just big enough to trap a single cell. This hook can be altered depending on the size and type of cells a researcher is using.
The first plunger is also used to wash and separate the captured cell from other cells that may have been extracted. The second plunger pushes the captured cell out of the pipette into growth medium or onto a slide or welled plate for study.
Dr Qin said one of his goals is to make the technology cost $10 or less per run. And future designs of the hSCP will be developed with mass production in mind.
Dr Qin said his group can also produce hSCPs that pick up virtually any small number of cells, depending on a scientist’s needs, by etching more hooks during the pipette’s construction. ![]()

Credit: Rhoda Baer
Researchers say they have developed a pipette that can transfer a single cell at a time.
The device, called the handheld single-cell pipette (hSCP), has 2 plungers. The first is used to wash and capture a single cell, and the second can release the cell in the desired location.
Lidong Qin, PhD, of Houston Methodist Research Institute in Texas, and his colleagues described the hSCP and reported preliminary results with the device in the Journal of the American Chemical Society.
“Studying single cells and their unique functions has become a frontier in current biomedical research,” Dr Qin said. “One of the biggest challenges for single-cell research is picking out only one cell from a collection of millions of cells.”
He noted that current techniques for withdrawing single cells from a tube or Petri dish can be cumbersome, expensive, and time-consuming.
“Some old and clumsy methods are used to capture single cells,” he explained. “Some researchers use their mouths at one end of the pipette, driven by their own mouth force, to try to ensure only a minimum amount of cell suspension collected. The sample is then checked with a microscope to find out the number of cells captured. The opportunity to get only one cell is hit-or-miss and a bit troublesome.”
“One company provides a million-dollar machine that can help biologists transfer single cells to 96-well plates. Each run costs an additional $1000 to purchase the plate. Such technology will not be widely accessible to biologists.”
With that in mind, Dr Qin and his colleagues developed their 2-plunger hSCP. The first plunger withdraws fluid from a suspension of cells.
Fluid travels through canals on either side of a nanoscopic, laser-sculpted “hook” that is just big enough to trap a single cell. This hook can be altered depending on the size and type of cells a researcher is using.
The first plunger is also used to wash and separate the captured cell from other cells that may have been extracted. The second plunger pushes the captured cell out of the pipette into growth medium or onto a slide or welled plate for study.
Dr Qin said one of his goals is to make the technology cost $10 or less per run. And future designs of the hSCP will be developed with mass production in mind.
Dr Qin said his group can also produce hSCPs that pick up virtually any small number of cells, depending on a scientist’s needs, by etching more hooks during the pipette’s construction. ![]()

Credit: Rhoda Baer
Researchers say they have developed a pipette that can transfer a single cell at a time.
The device, called the handheld single-cell pipette (hSCP), has 2 plungers. The first is used to wash and capture a single cell, and the second can release the cell in the desired location.
Lidong Qin, PhD, of Houston Methodist Research Institute in Texas, and his colleagues described the hSCP and reported preliminary results with the device in the Journal of the American Chemical Society.
“Studying single cells and their unique functions has become a frontier in current biomedical research,” Dr Qin said. “One of the biggest challenges for single-cell research is picking out only one cell from a collection of millions of cells.”
He noted that current techniques for withdrawing single cells from a tube or Petri dish can be cumbersome, expensive, and time-consuming.
“Some old and clumsy methods are used to capture single cells,” he explained. “Some researchers use their mouths at one end of the pipette, driven by their own mouth force, to try to ensure only a minimum amount of cell suspension collected. The sample is then checked with a microscope to find out the number of cells captured. The opportunity to get only one cell is hit-or-miss and a bit troublesome.”
“One company provides a million-dollar machine that can help biologists transfer single cells to 96-well plates. Each run costs an additional $1000 to purchase the plate. Such technology will not be widely accessible to biologists.”
With that in mind, Dr Qin and his colleagues developed their 2-plunger hSCP. The first plunger withdraws fluid from a suspension of cells.
Fluid travels through canals on either side of a nanoscopic, laser-sculpted “hook” that is just big enough to trap a single cell. This hook can be altered depending on the size and type of cells a researcher is using.
The first plunger is also used to wash and separate the captured cell from other cells that may have been extracted. The second plunger pushes the captured cell out of the pipette into growth medium or onto a slide or welled plate for study.
Dr Qin said one of his goals is to make the technology cost $10 or less per run. And future designs of the hSCP will be developed with mass production in mind.
Dr Qin said his group can also produce hSCPs that pick up virtually any small number of cells, depending on a scientist’s needs, by etching more hooks during the pipette’s construction. ![]()
Combo offers better detection of invasive aspergillosis
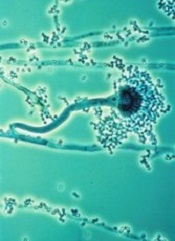
Results of a retrospective study may have revealed the most accurate way to diagnose invasive aspergillosis (IA).
The fungal infection can be life-threatening, particularly for immunosuppressed patients, but it remains difficult to diagnose.
So researchers compared 3 tests used to diagnose IA and found the combination of nucleic acid sequence-based amplification (NASBA) and real-time quantitative PCR (qPCR) had a 100% positive predictive value.
The team reported this discovery in The Journal of Molecular Diagnostics.
IA is caused by the fungus Aspergillus fumigatus, which is considered by many pathologists to be the world’s most harmful mold.
“Traditional diagnostic methods, such as culture and histopathology of infected tissues, often fail to detect Aspergillus,” said study investigator Yun Xia, PhD, of the First Affiliated Hospital of Chongqing Medical University in China.
With this in mind, he and his colleagues evaluated the diagnostic performance of 2 nucleic acid amplification assays—qPCR and NASBA—and 1 antigen-detection method—galactomannan enzyme-linked immunosorbent assay (GM-ELISA)—using blood samples from 80 patients at high risk of IA.
The researchers evaluated the tests alone and in combination. Of the 80 patients, 42.5% had proven or probable IA.
Tests showed that NASBA predicted IA with the highest sensitivity—76.47%, compared to 67.65% for qPCR and 52.94% for GM-ELISA. But qPCR offered the highest specificity—89.13%, compared to 80.43% for both NASBA and GM-ELISA.
NASBA had the highest negative predictive value—82.22%, compared to 78.85% for qPCR and 69.81% for GM-ELISA. And qPCR had the highest positive predictive value—82.14%, compared to 74.29% for NASBA and 66.67% for GM-ELISA.
NASBA and qPCR each had a high Youden index as well—0.5690 and 0.5678, respectively—compared to GM-ELISA—0.3337.
And combining the tests improved their accuracy. The combination of NASBA and qPCR led to 100% specificity and a 100% positive predictive value.
Dr Xia and his colleagues also noted that NASBA offers the advantages of rapid amplification (90 minutes) and simple operation with low instrument cost, compared with qPCR and GM-ELISA.
Finally, the team stressed that although GM-ELISA is widely and routinely used for aspergillosis diagnosis, this study indicates that it is inferior to both NASBA and qPCR. ![]()

Results of a retrospective study may have revealed the most accurate way to diagnose invasive aspergillosis (IA).
The fungal infection can be life-threatening, particularly for immunosuppressed patients, but it remains difficult to diagnose.
So researchers compared 3 tests used to diagnose IA and found the combination of nucleic acid sequence-based amplification (NASBA) and real-time quantitative PCR (qPCR) had a 100% positive predictive value.
The team reported this discovery in The Journal of Molecular Diagnostics.
IA is caused by the fungus Aspergillus fumigatus, which is considered by many pathologists to be the world’s most harmful mold.
“Traditional diagnostic methods, such as culture and histopathology of infected tissues, often fail to detect Aspergillus,” said study investigator Yun Xia, PhD, of the First Affiliated Hospital of Chongqing Medical University in China.
With this in mind, he and his colleagues evaluated the diagnostic performance of 2 nucleic acid amplification assays—qPCR and NASBA—and 1 antigen-detection method—galactomannan enzyme-linked immunosorbent assay (GM-ELISA)—using blood samples from 80 patients at high risk of IA.
The researchers evaluated the tests alone and in combination. Of the 80 patients, 42.5% had proven or probable IA.
Tests showed that NASBA predicted IA with the highest sensitivity—76.47%, compared to 67.65% for qPCR and 52.94% for GM-ELISA. But qPCR offered the highest specificity—89.13%, compared to 80.43% for both NASBA and GM-ELISA.
NASBA had the highest negative predictive value—82.22%, compared to 78.85% for qPCR and 69.81% for GM-ELISA. And qPCR had the highest positive predictive value—82.14%, compared to 74.29% for NASBA and 66.67% for GM-ELISA.
NASBA and qPCR each had a high Youden index as well—0.5690 and 0.5678, respectively—compared to GM-ELISA—0.3337.
And combining the tests improved their accuracy. The combination of NASBA and qPCR led to 100% specificity and a 100% positive predictive value.
Dr Xia and his colleagues also noted that NASBA offers the advantages of rapid amplification (90 minutes) and simple operation with low instrument cost, compared with qPCR and GM-ELISA.
Finally, the team stressed that although GM-ELISA is widely and routinely used for aspergillosis diagnosis, this study indicates that it is inferior to both NASBA and qPCR. ![]()

Results of a retrospective study may have revealed the most accurate way to diagnose invasive aspergillosis (IA).
The fungal infection can be life-threatening, particularly for immunosuppressed patients, but it remains difficult to diagnose.
So researchers compared 3 tests used to diagnose IA and found the combination of nucleic acid sequence-based amplification (NASBA) and real-time quantitative PCR (qPCR) had a 100% positive predictive value.
The team reported this discovery in The Journal of Molecular Diagnostics.
IA is caused by the fungus Aspergillus fumigatus, which is considered by many pathologists to be the world’s most harmful mold.
“Traditional diagnostic methods, such as culture and histopathology of infected tissues, often fail to detect Aspergillus,” said study investigator Yun Xia, PhD, of the First Affiliated Hospital of Chongqing Medical University in China.
With this in mind, he and his colleagues evaluated the diagnostic performance of 2 nucleic acid amplification assays—qPCR and NASBA—and 1 antigen-detection method—galactomannan enzyme-linked immunosorbent assay (GM-ELISA)—using blood samples from 80 patients at high risk of IA.
The researchers evaluated the tests alone and in combination. Of the 80 patients, 42.5% had proven or probable IA.
Tests showed that NASBA predicted IA with the highest sensitivity—76.47%, compared to 67.65% for qPCR and 52.94% for GM-ELISA. But qPCR offered the highest specificity—89.13%, compared to 80.43% for both NASBA and GM-ELISA.
NASBA had the highest negative predictive value—82.22%, compared to 78.85% for qPCR and 69.81% for GM-ELISA. And qPCR had the highest positive predictive value—82.14%, compared to 74.29% for NASBA and 66.67% for GM-ELISA.
NASBA and qPCR each had a high Youden index as well—0.5690 and 0.5678, respectively—compared to GM-ELISA—0.3337.
And combining the tests improved their accuracy. The combination of NASBA and qPCR led to 100% specificity and a 100% positive predictive value.
Dr Xia and his colleagues also noted that NASBA offers the advantages of rapid amplification (90 minutes) and simple operation with low instrument cost, compared with qPCR and GM-ELISA.
Finally, the team stressed that although GM-ELISA is widely and routinely used for aspergillosis diagnosis, this study indicates that it is inferior to both NASBA and qPCR. ![]()
Tool reveals how malaria parasites infect RBCs
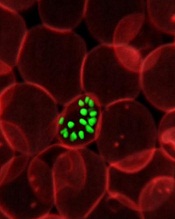
an RBC; Credit: St Jude
Children’s Research Hospital
Researchers say laser optical tweezers have allowed them to study how Plasmodium falciparum interacts with red blood cells (RBCs) at the single-cell level.
The research has revealed new insights into malaria biology and may pave the way for more effective drugs or vaccines.
Julian Rayner, PhD, of the Wellcome Trust Sanger Institute in Cambridge, UK, and his colleagues described their use of laser optical tweezers in Biophysical Journal.
“Using laser tweezers to study red blood cell invasion gives us an unprecedented level of control over the whole process and will help us to understand this critical process at a level of detail that has not been possible before,” Dr Rayner said.
He and his colleagues noted that P falciparum merozoites usually leave one RBC and invade another in less than a minute. And the merozoites lose the ability to infect host cells within 2 or 3 minutes of release.
So the researchers used laser optical tweezers to study this transient event. The tweezers allow for precise control over the movements of cells by exerting extremely small forces with a highly focused laser beam.
The team used the tweezers to pick up individual merozoites that had just emerged from an RBC and deliver them to another RBC, demonstrating that the technique is suitable for studying the invasion process.
The researchers also used the tweezers to measure how strongly the merozoites adhere to RBCs. They discovered that attachment is probably mediated by multiple weak interactions, which could potentially be blocked by a combination of drugs or antibodies.
Finally, the team used the tweezers to shed light on how 3 different invasion-inhibiting drugs—heparin, cytochalasin D, and chymotrypsin—affect interactions between merozoites and RBCs.
The tweezers revealed that heparin blocks merozoite attachment to any surface, including glass slides. This suggests a receptor-independent mode of action, which contradicts the previously proposed mechanism.
Cytochalasin D, on the other hand, had no effect on attachment force, a finding that also contradicts previous thought.
And with chymotrypsin, the researchers observed 2 different effects. When merozoites adhered to chymotrypsin-treated RBCs, they did so with a reduction in the force of attachment that was similar to the effect the enzyme had on the overall efficiency of invasion.
However, merozoites that had been released more than 3 minutes previously were no longer able to adhere to chymotrypsin-treated RBCs. This suggests that chymotrypsin affects both the force of merozoite attachment and the time in which invasion can occur.
Taken together, these findings show that optical tweezers enable the study of malaria biology and drug mechanisms at the single-cell level.
“We now plan to apply this technology to dissect the process of invasion and understand what genes and proteins function at what step,” Dr Rayner said. “This will allow us to design better inhibitors or vaccines that block invasion by targeting multiple steps at the same time.” ![]()

an RBC; Credit: St Jude
Children’s Research Hospital
Researchers say laser optical tweezers have allowed them to study how Plasmodium falciparum interacts with red blood cells (RBCs) at the single-cell level.
The research has revealed new insights into malaria biology and may pave the way for more effective drugs or vaccines.
Julian Rayner, PhD, of the Wellcome Trust Sanger Institute in Cambridge, UK, and his colleagues described their use of laser optical tweezers in Biophysical Journal.
“Using laser tweezers to study red blood cell invasion gives us an unprecedented level of control over the whole process and will help us to understand this critical process at a level of detail that has not been possible before,” Dr Rayner said.
He and his colleagues noted that P falciparum merozoites usually leave one RBC and invade another in less than a minute. And the merozoites lose the ability to infect host cells within 2 or 3 minutes of release.
So the researchers used laser optical tweezers to study this transient event. The tweezers allow for precise control over the movements of cells by exerting extremely small forces with a highly focused laser beam.
The team used the tweezers to pick up individual merozoites that had just emerged from an RBC and deliver them to another RBC, demonstrating that the technique is suitable for studying the invasion process.
The researchers also used the tweezers to measure how strongly the merozoites adhere to RBCs. They discovered that attachment is probably mediated by multiple weak interactions, which could potentially be blocked by a combination of drugs or antibodies.
Finally, the team used the tweezers to shed light on how 3 different invasion-inhibiting drugs—heparin, cytochalasin D, and chymotrypsin—affect interactions between merozoites and RBCs.
The tweezers revealed that heparin blocks merozoite attachment to any surface, including glass slides. This suggests a receptor-independent mode of action, which contradicts the previously proposed mechanism.
Cytochalasin D, on the other hand, had no effect on attachment force, a finding that also contradicts previous thought.
And with chymotrypsin, the researchers observed 2 different effects. When merozoites adhered to chymotrypsin-treated RBCs, they did so with a reduction in the force of attachment that was similar to the effect the enzyme had on the overall efficiency of invasion.
However, merozoites that had been released more than 3 minutes previously were no longer able to adhere to chymotrypsin-treated RBCs. This suggests that chymotrypsin affects both the force of merozoite attachment and the time in which invasion can occur.
Taken together, these findings show that optical tweezers enable the study of malaria biology and drug mechanisms at the single-cell level.
“We now plan to apply this technology to dissect the process of invasion and understand what genes and proteins function at what step,” Dr Rayner said. “This will allow us to design better inhibitors or vaccines that block invasion by targeting multiple steps at the same time.” ![]()

an RBC; Credit: St Jude
Children’s Research Hospital
Researchers say laser optical tweezers have allowed them to study how Plasmodium falciparum interacts with red blood cells (RBCs) at the single-cell level.
The research has revealed new insights into malaria biology and may pave the way for more effective drugs or vaccines.
Julian Rayner, PhD, of the Wellcome Trust Sanger Institute in Cambridge, UK, and his colleagues described their use of laser optical tweezers in Biophysical Journal.
“Using laser tweezers to study red blood cell invasion gives us an unprecedented level of control over the whole process and will help us to understand this critical process at a level of detail that has not been possible before,” Dr Rayner said.
He and his colleagues noted that P falciparum merozoites usually leave one RBC and invade another in less than a minute. And the merozoites lose the ability to infect host cells within 2 or 3 minutes of release.
So the researchers used laser optical tweezers to study this transient event. The tweezers allow for precise control over the movements of cells by exerting extremely small forces with a highly focused laser beam.
The team used the tweezers to pick up individual merozoites that had just emerged from an RBC and deliver them to another RBC, demonstrating that the technique is suitable for studying the invasion process.
The researchers also used the tweezers to measure how strongly the merozoites adhere to RBCs. They discovered that attachment is probably mediated by multiple weak interactions, which could potentially be blocked by a combination of drugs or antibodies.
Finally, the team used the tweezers to shed light on how 3 different invasion-inhibiting drugs—heparin, cytochalasin D, and chymotrypsin—affect interactions between merozoites and RBCs.
The tweezers revealed that heparin blocks merozoite attachment to any surface, including glass slides. This suggests a receptor-independent mode of action, which contradicts the previously proposed mechanism.
Cytochalasin D, on the other hand, had no effect on attachment force, a finding that also contradicts previous thought.
And with chymotrypsin, the researchers observed 2 different effects. When merozoites adhered to chymotrypsin-treated RBCs, they did so with a reduction in the force of attachment that was similar to the effect the enzyme had on the overall efficiency of invasion.
However, merozoites that had been released more than 3 minutes previously were no longer able to adhere to chymotrypsin-treated RBCs. This suggests that chymotrypsin affects both the force of merozoite attachment and the time in which invasion can occur.
Taken together, these findings show that optical tweezers enable the study of malaria biology and drug mechanisms at the single-cell level.
“We now plan to apply this technology to dissect the process of invasion and understand what genes and proteins function at what step,” Dr Rayner said. “This will allow us to design better inhibitors or vaccines that block invasion by targeting multiple steps at the same time.”
Discovery could help halt malaria transmission

Credit: Swiss TPH
Malaria parasites exploit the epigenetic regulator HP1 to promote their survival and transmission between human hosts, a new study suggests.
It appears that Plasmodium falciparum uses HP1 to control the expression of surface antigens and escape the body’s immune responses. This prolongs the parasite’s survival and enables its transmission.
Researchers believe this discovery paves the way for new strategies to prevent malaria transmission.
Till Voss, PhD, of the Swiss Tropical and Public Health Institute in Basel, and his colleagues detailed the discovery in Cell Host & Microbe.
The team knew that HP1 induces heritable condensation of chromosomal regions. As a result, genes located within these regions are not expressed.
Since this conformation is reversible, HP1-controlled genes can become activated without requiring changes in the underlying DNA sequence.
With this in mind, the researchers engineered a mutant P falciparum parasite in which HP1 expression can be shut down. And the team observed that, in HP1-depleted parasites, all of the 60 var genes became highly active.
Each var gene encodes a distinct variant of the virulence factor PfEMP1, which is displayed on the surface of the parasite-infected red blood cell. PfEMP1 is a major target of the immune system in infected humans.
Individual parasites normally express only 1 of the 60 var/PfEMP1 proteins, while keeping all other members silenced. By switching to another var/PfEMP1 variant, the parasite is able to escape existing immune responses raised against previous variants.
Dr Voss and his colleagues found that HP1 protects the PfEMP1 antigenic repertoire from being exposed to the immune system all at once.
“This finding is a major step forward in understanding the complex mechanisms responsible for antigenic variation,” Dr Voss said. “Furthermore, the tools generated in our study may be relevant for future research on malaria vaccines and immunity.”
The researchers also found that parasites lacking HP1 fail to copy their genomes and are therefore unable to proliferate. Initially, this led the team to believe that all the parasites they had cultured were dead.
However, more than 50% of these parasites turned out to be fully viable and differentiated into gametocytes, the sexual form of the malaria parasite. Gametocytes are the only form of the parasite capable of infecting a mosquito and are a prerequisite to transmit malaria between humans.
“Such a high sexual conversion rate is unprecedented,” Dr Voss said. “Usually, only around 1% of parasites undergo this switch.”
Further experiments revealed that a master transcription factor triggering sexual differentiation—AP2-G—is expressed at much higher levels in parasites lacking HP1. Under normal conditions, HP1 silences the expression of AP2-G and therefore prevents sexual conversion in most parasites.
“The switch from parasite proliferation to gametocyte differentiation is controlled epigenetically by an HP1-dependent mechanism,” Dr Voss said.
“With this knowledge in hand, and with the identification of another epigenetic regulator involved in the same process [also published in Cell Host & Microbe], we are now able to specifically track the sexual conversion pathway in molecular detail.”
This may enable the development of new drugs to prevent sexual conversion and, consequently, malaria transmission.

Credit: Swiss TPH
Malaria parasites exploit the epigenetic regulator HP1 to promote their survival and transmission between human hosts, a new study suggests.
It appears that Plasmodium falciparum uses HP1 to control the expression of surface antigens and escape the body’s immune responses. This prolongs the parasite’s survival and enables its transmission.
Researchers believe this discovery paves the way for new strategies to prevent malaria transmission.
Till Voss, PhD, of the Swiss Tropical and Public Health Institute in Basel, and his colleagues detailed the discovery in Cell Host & Microbe.
The team knew that HP1 induces heritable condensation of chromosomal regions. As a result, genes located within these regions are not expressed.
Since this conformation is reversible, HP1-controlled genes can become activated without requiring changes in the underlying DNA sequence.
With this in mind, the researchers engineered a mutant P falciparum parasite in which HP1 expression can be shut down. And the team observed that, in HP1-depleted parasites, all of the 60 var genes became highly active.
Each var gene encodes a distinct variant of the virulence factor PfEMP1, which is displayed on the surface of the parasite-infected red blood cell. PfEMP1 is a major target of the immune system in infected humans.
Individual parasites normally express only 1 of the 60 var/PfEMP1 proteins, while keeping all other members silenced. By switching to another var/PfEMP1 variant, the parasite is able to escape existing immune responses raised against previous variants.
Dr Voss and his colleagues found that HP1 protects the PfEMP1 antigenic repertoire from being exposed to the immune system all at once.
“This finding is a major step forward in understanding the complex mechanisms responsible for antigenic variation,” Dr Voss said. “Furthermore, the tools generated in our study may be relevant for future research on malaria vaccines and immunity.”
The researchers also found that parasites lacking HP1 fail to copy their genomes and are therefore unable to proliferate. Initially, this led the team to believe that all the parasites they had cultured were dead.
However, more than 50% of these parasites turned out to be fully viable and differentiated into gametocytes, the sexual form of the malaria parasite. Gametocytes are the only form of the parasite capable of infecting a mosquito and are a prerequisite to transmit malaria between humans.
“Such a high sexual conversion rate is unprecedented,” Dr Voss said. “Usually, only around 1% of parasites undergo this switch.”
Further experiments revealed that a master transcription factor triggering sexual differentiation—AP2-G—is expressed at much higher levels in parasites lacking HP1. Under normal conditions, HP1 silences the expression of AP2-G and therefore prevents sexual conversion in most parasites.
“The switch from parasite proliferation to gametocyte differentiation is controlled epigenetically by an HP1-dependent mechanism,” Dr Voss said.
“With this knowledge in hand, and with the identification of another epigenetic regulator involved in the same process [also published in Cell Host & Microbe], we are now able to specifically track the sexual conversion pathway in molecular detail.”
This may enable the development of new drugs to prevent sexual conversion and, consequently, malaria transmission.

Credit: Swiss TPH
Malaria parasites exploit the epigenetic regulator HP1 to promote their survival and transmission between human hosts, a new study suggests.
It appears that Plasmodium falciparum uses HP1 to control the expression of surface antigens and escape the body’s immune responses. This prolongs the parasite’s survival and enables its transmission.
Researchers believe this discovery paves the way for new strategies to prevent malaria transmission.
Till Voss, PhD, of the Swiss Tropical and Public Health Institute in Basel, and his colleagues detailed the discovery in Cell Host & Microbe.
The team knew that HP1 induces heritable condensation of chromosomal regions. As a result, genes located within these regions are not expressed.
Since this conformation is reversible, HP1-controlled genes can become activated without requiring changes in the underlying DNA sequence.
With this in mind, the researchers engineered a mutant P falciparum parasite in which HP1 expression can be shut down. And the team observed that, in HP1-depleted parasites, all of the 60 var genes became highly active.
Each var gene encodes a distinct variant of the virulence factor PfEMP1, which is displayed on the surface of the parasite-infected red blood cell. PfEMP1 is a major target of the immune system in infected humans.
Individual parasites normally express only 1 of the 60 var/PfEMP1 proteins, while keeping all other members silenced. By switching to another var/PfEMP1 variant, the parasite is able to escape existing immune responses raised against previous variants.
Dr Voss and his colleagues found that HP1 protects the PfEMP1 antigenic repertoire from being exposed to the immune system all at once.
“This finding is a major step forward in understanding the complex mechanisms responsible for antigenic variation,” Dr Voss said. “Furthermore, the tools generated in our study may be relevant for future research on malaria vaccines and immunity.”
The researchers also found that parasites lacking HP1 fail to copy their genomes and are therefore unable to proliferate. Initially, this led the team to believe that all the parasites they had cultured were dead.
However, more than 50% of these parasites turned out to be fully viable and differentiated into gametocytes, the sexual form of the malaria parasite. Gametocytes are the only form of the parasite capable of infecting a mosquito and are a prerequisite to transmit malaria between humans.
“Such a high sexual conversion rate is unprecedented,” Dr Voss said. “Usually, only around 1% of parasites undergo this switch.”
Further experiments revealed that a master transcription factor triggering sexual differentiation—AP2-G—is expressed at much higher levels in parasites lacking HP1. Under normal conditions, HP1 silences the expression of AP2-G and therefore prevents sexual conversion in most parasites.
“The switch from parasite proliferation to gametocyte differentiation is controlled epigenetically by an HP1-dependent mechanism,” Dr Voss said.
“With this knowledge in hand, and with the identification of another epigenetic regulator involved in the same process [also published in Cell Host & Microbe], we are now able to specifically track the sexual conversion pathway in molecular detail.”
This may enable the development of new drugs to prevent sexual conversion and, consequently, malaria transmission.
A quicker way to manipulate malaria genes

(left) and Jeffrey Wagner
Credit: Bryce Vickmark
The gene-editing technique CRISPR can disrupt a single gene from the malaria parasite Plasmodium falciparum in a matter of weeks, a new study suggests.
Although CRISPR’s success rate ranged from 50% to 100%, the researchers believe the technique shows promise and could greatly speed up gene analysis.
At present, it can take up to a year to determine the function of a single gene in P falciparum, which can hinder efforts to develop drugs and vaccines.
“Even though we’ve sequenced the entire genome of Plasmodium falciparum, half of it still remains functionally uncharacterized,” said Jacquin Niles, MD, PhD, of the Massachusetts Institute of Technology in Cambridge.
“That’s about 2500 genes that, if only we knew what they did, we could think about novel therapeutics, whether it’s drugs or vaccines.”
Dr Niles and his colleagues described their use of CRISPR in P falciparum in Nature Methods.
The team noted that, in P falciparum, gene editing can take up to a year because it relies on homologous recombination, a type of genetic swapping that cells use to repair broken DNA strands and that occurs very rarely in the genome of the malaria parasite.
“You have to rely on this really inefficient process that occurs only if you have spontaneous DNA strand breaks that happen to fall within your region of interest,” Dr Niles said.
More recently, researchers have successfully used zinc finger nucleases to cut out specific genes, but this approach is costly because it requires a new nuclease to be designed for each gene target.
CRISPR exploits a set of bacterial proteins that protect microbes from viral infection. The system includes a DNA-cutting enzyme, Cas9, bound to a short RNA guide strand that is programmed to bind to a specific genome sequence, telling Cas9 where to make its cut. This approach allows scientists to target and delete any gene by simply changing the RNA guide strand sequence.
As soon as researchers proved this system could work in cells other than bacteria, Dr Niles started to think about using it to manipulate P falciparum.
To test this approach, he and his colleagues tried using CRISPR to disrupt 2 genes, kahrp and eba-175, that had previously been knocked out in malaria using traditional approaches.
The kahrp gene produces a protein that causes red blood cells to develop a knobby appearance when infected with malaria. Dr Niles’s team was able to disrupt this gene in 100% of parasites treated with the CRISPR system. The red blood cells infected by parasites remained smooth.
The other gene, eba-175, codes for a protein that binds to red blood cell receptors and helps the malaria parasite enter cells. The researchers were only able to disrupt this gene in 50% to 80% of parasites manipulated with CRISPR.
“We consider this to be a win,” Dr Niles said. “Compared to the efficiency with which P falciparum genetics have been done in the past, even 50% is pretty substantial.”
Now that CRISPR technology has been validated in P falciparum, Dr Niles expects many scientists will adopt it for genetic studies of the parasite. Such efforts could reveal more about how the parasite invades red blood cells and replicates inside cells, which could generate new drug and vaccine targets.
“I think the impact could be quite huge,” he said. “It lowers the barrier to really being more imaginative in terms of how we do experiments and the kinds of questions that we can ask.”

(left) and Jeffrey Wagner
Credit: Bryce Vickmark
The gene-editing technique CRISPR can disrupt a single gene from the malaria parasite Plasmodium falciparum in a matter of weeks, a new study suggests.
Although CRISPR’s success rate ranged from 50% to 100%, the researchers believe the technique shows promise and could greatly speed up gene analysis.
At present, it can take up to a year to determine the function of a single gene in P falciparum, which can hinder efforts to develop drugs and vaccines.
“Even though we’ve sequenced the entire genome of Plasmodium falciparum, half of it still remains functionally uncharacterized,” said Jacquin Niles, MD, PhD, of the Massachusetts Institute of Technology in Cambridge.
“That’s about 2500 genes that, if only we knew what they did, we could think about novel therapeutics, whether it’s drugs or vaccines.”
Dr Niles and his colleagues described their use of CRISPR in P falciparum in Nature Methods.
The team noted that, in P falciparum, gene editing can take up to a year because it relies on homologous recombination, a type of genetic swapping that cells use to repair broken DNA strands and that occurs very rarely in the genome of the malaria parasite.
“You have to rely on this really inefficient process that occurs only if you have spontaneous DNA strand breaks that happen to fall within your region of interest,” Dr Niles said.
More recently, researchers have successfully used zinc finger nucleases to cut out specific genes, but this approach is costly because it requires a new nuclease to be designed for each gene target.
CRISPR exploits a set of bacterial proteins that protect microbes from viral infection. The system includes a DNA-cutting enzyme, Cas9, bound to a short RNA guide strand that is programmed to bind to a specific genome sequence, telling Cas9 where to make its cut. This approach allows scientists to target and delete any gene by simply changing the RNA guide strand sequence.
As soon as researchers proved this system could work in cells other than bacteria, Dr Niles started to think about using it to manipulate P falciparum.
To test this approach, he and his colleagues tried using CRISPR to disrupt 2 genes, kahrp and eba-175, that had previously been knocked out in malaria using traditional approaches.
The kahrp gene produces a protein that causes red blood cells to develop a knobby appearance when infected with malaria. Dr Niles’s team was able to disrupt this gene in 100% of parasites treated with the CRISPR system. The red blood cells infected by parasites remained smooth.
The other gene, eba-175, codes for a protein that binds to red blood cell receptors and helps the malaria parasite enter cells. The researchers were only able to disrupt this gene in 50% to 80% of parasites manipulated with CRISPR.
“We consider this to be a win,” Dr Niles said. “Compared to the efficiency with which P falciparum genetics have been done in the past, even 50% is pretty substantial.”
Now that CRISPR technology has been validated in P falciparum, Dr Niles expects many scientists will adopt it for genetic studies of the parasite. Such efforts could reveal more about how the parasite invades red blood cells and replicates inside cells, which could generate new drug and vaccine targets.
“I think the impact could be quite huge,” he said. “It lowers the barrier to really being more imaginative in terms of how we do experiments and the kinds of questions that we can ask.”

(left) and Jeffrey Wagner
Credit: Bryce Vickmark
The gene-editing technique CRISPR can disrupt a single gene from the malaria parasite Plasmodium falciparum in a matter of weeks, a new study suggests.
Although CRISPR’s success rate ranged from 50% to 100%, the researchers believe the technique shows promise and could greatly speed up gene analysis.
At present, it can take up to a year to determine the function of a single gene in P falciparum, which can hinder efforts to develop drugs and vaccines.
“Even though we’ve sequenced the entire genome of Plasmodium falciparum, half of it still remains functionally uncharacterized,” said Jacquin Niles, MD, PhD, of the Massachusetts Institute of Technology in Cambridge.
“That’s about 2500 genes that, if only we knew what they did, we could think about novel therapeutics, whether it’s drugs or vaccines.”
Dr Niles and his colleagues described their use of CRISPR in P falciparum in Nature Methods.
The team noted that, in P falciparum, gene editing can take up to a year because it relies on homologous recombination, a type of genetic swapping that cells use to repair broken DNA strands and that occurs very rarely in the genome of the malaria parasite.
“You have to rely on this really inefficient process that occurs only if you have spontaneous DNA strand breaks that happen to fall within your region of interest,” Dr Niles said.
More recently, researchers have successfully used zinc finger nucleases to cut out specific genes, but this approach is costly because it requires a new nuclease to be designed for each gene target.
CRISPR exploits a set of bacterial proteins that protect microbes from viral infection. The system includes a DNA-cutting enzyme, Cas9, bound to a short RNA guide strand that is programmed to bind to a specific genome sequence, telling Cas9 where to make its cut. This approach allows scientists to target and delete any gene by simply changing the RNA guide strand sequence.
As soon as researchers proved this system could work in cells other than bacteria, Dr Niles started to think about using it to manipulate P falciparum.
To test this approach, he and his colleagues tried using CRISPR to disrupt 2 genes, kahrp and eba-175, that had previously been knocked out in malaria using traditional approaches.
The kahrp gene produces a protein that causes red blood cells to develop a knobby appearance when infected with malaria. Dr Niles’s team was able to disrupt this gene in 100% of parasites treated with the CRISPR system. The red blood cells infected by parasites remained smooth.
The other gene, eba-175, codes for a protein that binds to red blood cell receptors and helps the malaria parasite enter cells. The researchers were only able to disrupt this gene in 50% to 80% of parasites manipulated with CRISPR.
“We consider this to be a win,” Dr Niles said. “Compared to the efficiency with which P falciparum genetics have been done in the past, even 50% is pretty substantial.”
Now that CRISPR technology has been validated in P falciparum, Dr Niles expects many scientists will adopt it for genetic studies of the parasite. Such efforts could reveal more about how the parasite invades red blood cells and replicates inside cells, which could generate new drug and vaccine targets.
“I think the impact could be quite huge,” he said. “It lowers the barrier to really being more imaginative in terms of how we do experiments and the kinds of questions that we can ask.”
Viruses can protect mice from malaria

Credit: Ute Frevert
and Margaret Shear
In a new study, genetically altered viruses produced long-lasting antimalaria antibodies in mice and protected many of them from the disease.
The approach, known as vector immunoprophylaxis (VIP), produced antibodies against the Plasmodium falciparum circumsporozoite protein (CSP) and prevented malaria infection in 10% to 100% of mice, depending on the dose and type of viral vector used.
Researchers recounted these results in PNAS.
“We need better ways to fight malaria,” said study author Gary Ketner, PhD, of the Johns Hopkins Bloomberg School of Public Health in Baltimore, Maryland. “And our research suggests [VIP] could be a promising approach.”
To test the approach, Dr Ketner and his colleagues constructed adeno-associated virus (AAV) vectors encoding human immunoglobulin G (hIgG) specific for the P falciparum CSP central repeat by inserting the variable regions of mouse monoclonal antibodies (mAbs) 2A10 and 2C11 into the hIgG framework of the VIP expression plasmid.
The team then injected mice with 1 x 1011 genome copies (GC) of 2A10-AAV, 2C11-AAV, b12-AAV (which protects against HIV), or with buffer.
Within a week, the AAV-transduced mice expressed hIgG at 50 μg/mL to 500 μg/mL in serum. The expression increased until about the 4-week mark, when it reached 1000 μg/mL in some mice.
The mice that received 2A10-AAV or 2C11-AAV expressed antibodies that bound recombinant CSP and recognized whole P falciparum sporozoites. The b12-AAV-transduced mice and buffer-transduced mice did not.
In all AAV-transduced mice, serum antibody concentrations plateaued at 4 to 8 weeks and remained at that level through the end of the study, which was 52 weeks after transduction.
At the 8-week mark, the researchers tested the efficacy of VIP. They introduced—either intravenously or through a mosquito-bite challenge—transgenic Plasmodium berghei rodent sporozoites that incorporate the P falciparum target of the antibody in their CSP.
In the intravenously challenged group, 70% of 2A10-AAV-transduced mice were protected from malaria. In the mosquito-bite-challenged group, 60% of 2A10-AAV-transduced mice and 30% of 2C11-AAV-transduced mice were protected from malaria.
Role of dose and antibody level
To examine the effects of vector dose on mAb production and malaria protection, the researchers compared varying doses of 2A10-AAV to b12-AAV. They tested mice transduced with 3 x 1011 GC of b12-AAV or doses of 2A10-AAV ranging from 3 x 109 GC to 3 x 1011 GC.
The team conducted a mosquito-bite challenge at 11 weeks after transduction. And they found that 70% of the mice that received the highest AAV dose (1 x 1011 GC) were protected, as were 40% of mice that received 3 x 1010 GC and 10% of mice that received 1 x 1010 GC.
All mice transduced with 3 x 109 GC were parasitemic by day 7, and all b12-AAV mice were parasitemic by day 6. There was a signficant correlation between 2A10 antibody concentration and day to parasitemia.
In a subset of mice that produced higher levels of antibodies, the antimalaria protection was 100%. These mice expressed CSP-specific mAb 2A10 at 1 mg/mL or more.
So it seems the protection from malaria is dose-dependent, said study author Cailin Deal, PhD, of the Ragon Institute of MGH, MIT and Harvard in Cambridge, Massachusetts.
“Of course, we don’t know what the human dosage would be,” she added, “but it’s conceivable that the right dosage could completely protect against malaria.”

Credit: Ute Frevert
and Margaret Shear
In a new study, genetically altered viruses produced long-lasting antimalaria antibodies in mice and protected many of them from the disease.
The approach, known as vector immunoprophylaxis (VIP), produced antibodies against the Plasmodium falciparum circumsporozoite protein (CSP) and prevented malaria infection in 10% to 100% of mice, depending on the dose and type of viral vector used.
Researchers recounted these results in PNAS.
“We need better ways to fight malaria,” said study author Gary Ketner, PhD, of the Johns Hopkins Bloomberg School of Public Health in Baltimore, Maryland. “And our research suggests [VIP] could be a promising approach.”
To test the approach, Dr Ketner and his colleagues constructed adeno-associated virus (AAV) vectors encoding human immunoglobulin G (hIgG) specific for the P falciparum CSP central repeat by inserting the variable regions of mouse monoclonal antibodies (mAbs) 2A10 and 2C11 into the hIgG framework of the VIP expression plasmid.
The team then injected mice with 1 x 1011 genome copies (GC) of 2A10-AAV, 2C11-AAV, b12-AAV (which protects against HIV), or with buffer.
Within a week, the AAV-transduced mice expressed hIgG at 50 μg/mL to 500 μg/mL in serum. The expression increased until about the 4-week mark, when it reached 1000 μg/mL in some mice.
The mice that received 2A10-AAV or 2C11-AAV expressed antibodies that bound recombinant CSP and recognized whole P falciparum sporozoites. The b12-AAV-transduced mice and buffer-transduced mice did not.
In all AAV-transduced mice, serum antibody concentrations plateaued at 4 to 8 weeks and remained at that level through the end of the study, which was 52 weeks after transduction.
At the 8-week mark, the researchers tested the efficacy of VIP. They introduced—either intravenously or through a mosquito-bite challenge—transgenic Plasmodium berghei rodent sporozoites that incorporate the P falciparum target of the antibody in their CSP.
In the intravenously challenged group, 70% of 2A10-AAV-transduced mice were protected from malaria. In the mosquito-bite-challenged group, 60% of 2A10-AAV-transduced mice and 30% of 2C11-AAV-transduced mice were protected from malaria.
Role of dose and antibody level
To examine the effects of vector dose on mAb production and malaria protection, the researchers compared varying doses of 2A10-AAV to b12-AAV. They tested mice transduced with 3 x 1011 GC of b12-AAV or doses of 2A10-AAV ranging from 3 x 109 GC to 3 x 1011 GC.
The team conducted a mosquito-bite challenge at 11 weeks after transduction. And they found that 70% of the mice that received the highest AAV dose (1 x 1011 GC) were protected, as were 40% of mice that received 3 x 1010 GC and 10% of mice that received 1 x 1010 GC.
All mice transduced with 3 x 109 GC were parasitemic by day 7, and all b12-AAV mice were parasitemic by day 6. There was a signficant correlation between 2A10 antibody concentration and day to parasitemia.
In a subset of mice that produced higher levels of antibodies, the antimalaria protection was 100%. These mice expressed CSP-specific mAb 2A10 at 1 mg/mL or more.
So it seems the protection from malaria is dose-dependent, said study author Cailin Deal, PhD, of the Ragon Institute of MGH, MIT and Harvard in Cambridge, Massachusetts.
“Of course, we don’t know what the human dosage would be,” she added, “but it’s conceivable that the right dosage could completely protect against malaria.”

Credit: Ute Frevert
and Margaret Shear
In a new study, genetically altered viruses produced long-lasting antimalaria antibodies in mice and protected many of them from the disease.
The approach, known as vector immunoprophylaxis (VIP), produced antibodies against the Plasmodium falciparum circumsporozoite protein (CSP) and prevented malaria infection in 10% to 100% of mice, depending on the dose and type of viral vector used.
Researchers recounted these results in PNAS.
“We need better ways to fight malaria,” said study author Gary Ketner, PhD, of the Johns Hopkins Bloomberg School of Public Health in Baltimore, Maryland. “And our research suggests [VIP] could be a promising approach.”
To test the approach, Dr Ketner and his colleagues constructed adeno-associated virus (AAV) vectors encoding human immunoglobulin G (hIgG) specific for the P falciparum CSP central repeat by inserting the variable regions of mouse monoclonal antibodies (mAbs) 2A10 and 2C11 into the hIgG framework of the VIP expression plasmid.
The team then injected mice with 1 x 1011 genome copies (GC) of 2A10-AAV, 2C11-AAV, b12-AAV (which protects against HIV), or with buffer.
Within a week, the AAV-transduced mice expressed hIgG at 50 μg/mL to 500 μg/mL in serum. The expression increased until about the 4-week mark, when it reached 1000 μg/mL in some mice.
The mice that received 2A10-AAV or 2C11-AAV expressed antibodies that bound recombinant CSP and recognized whole P falciparum sporozoites. The b12-AAV-transduced mice and buffer-transduced mice did not.
In all AAV-transduced mice, serum antibody concentrations plateaued at 4 to 8 weeks and remained at that level through the end of the study, which was 52 weeks after transduction.
At the 8-week mark, the researchers tested the efficacy of VIP. They introduced—either intravenously or through a mosquito-bite challenge—transgenic Plasmodium berghei rodent sporozoites that incorporate the P falciparum target of the antibody in their CSP.
In the intravenously challenged group, 70% of 2A10-AAV-transduced mice were protected from malaria. In the mosquito-bite-challenged group, 60% of 2A10-AAV-transduced mice and 30% of 2C11-AAV-transduced mice were protected from malaria.
Role of dose and antibody level
To examine the effects of vector dose on mAb production and malaria protection, the researchers compared varying doses of 2A10-AAV to b12-AAV. They tested mice transduced with 3 x 1011 GC of b12-AAV or doses of 2A10-AAV ranging from 3 x 109 GC to 3 x 1011 GC.
The team conducted a mosquito-bite challenge at 11 weeks after transduction. And they found that 70% of the mice that received the highest AAV dose (1 x 1011 GC) were protected, as were 40% of mice that received 3 x 1010 GC and 10% of mice that received 1 x 1010 GC.
All mice transduced with 3 x 109 GC were parasitemic by day 7, and all b12-AAV mice were parasitemic by day 6. There was a signficant correlation between 2A10 antibody concentration and day to parasitemia.
In a subset of mice that produced higher levels of antibodies, the antimalaria protection was 100%. These mice expressed CSP-specific mAb 2A10 at 1 mg/mL or more.
So it seems the protection from malaria is dose-dependent, said study author Cailin Deal, PhD, of the Ragon Institute of MGH, MIT and Harvard in Cambridge, Massachusetts.
“Of course, we don’t know what the human dosage would be,” she added, “but it’s conceivable that the right dosage could completely protect against malaria.”