User login
Team reprograms blood cells into HSCs in mice

Researchers have found a way to reprogram mature blood cells from mice into hematopoietic stem cells (HSCs), according to a paper published in Cell.
The team used 8 transcription factors to reprogram blood progenitor cells and mature mouse myeloid cells into HSCs.
These cells, called induced HSCs (iHSCs), have the functional hallmarks of natural HSCs, are able to self-renew like natural HSCs, and can give rise to all of the cellular components of the blood.
“Blood cell production invariably goes in one direction—from stem cells, to progenitors, to mature effector cells,” said study author Derrick J. Rossi, PhD, of Boston Children’s Hospital in Massachusetts.
“We wanted to reverse the process and derive HSCs from differentiated blood cells using transcription factors that we found were specific to HSCs.”
To that end, Dr Rossi and his colleagues screened gene expression in 40 different types of blood and blood progenitor cells from mice. From this screen, the team identified 36 transcription factors that are expressed in HSCs but not in the cells that arise from them.
In a series of mouse transplantation experiments, the researchers found that 6 of the 36 transcription factors—Hlf, Runx1t1, Pbx1, Lmo2, Zfp37, and Prdm5—plus 2 additional factors not originally identified in their screen—Mycn and Meis1—were sufficient to reprogram 2 kinds of blood progenitor cells—pro/pre-B cells and common myeloid progenitor cells—into iHSCs.
The team reprogrammed their source cells by exposing them to viruses containing the genes for all 8 transcription factors and a molecular switch that turned the factor genes on in the presence of doxycycline. They then transplanted the exposed cells into recipient mice and activated the genes by giving the mice doxycycline.
The resulting iHSCs were capable of generating the entire blood cell repertoire in the transplanted mice, showing they had gained the ability to differentiate into all blood lineages. Stem cells collected from those recipients were capable of reconstituting the blood of secondary transplant recipients, proving that the 8-factor cocktail could instill the capacity for self-renewal.
Taking the work a step further, the researchers treated mature mouse myeloid cells with the same 8-factor cocktail. The resulting iHSCs produced all of the blood lineages and could regenerate the blood of secondary transplant recipients.
Study author Stuart Orkin, MD, of the Dana-Farber Cancer Institute in Boston, noted that the use of mice as a kind of reactor for reprogramming marks a novel direction in HSC research.
“In the blood research field, no one has the conditions to expand HSCs in the tissue culture dish,” he said. “Instead, by letting the reprogramming occur in mice, Rossi takes advantage of the signaling and environmental cues HSCs would normally experience.”
Dr Orkin added that iHSCs are nearly indistinguishable from normal HSCs at the transcriptional level. Unfortunately, though, these findings are far from translation to the clinic.
Researchers must still ascertain the precise contribution each of the 8 transcription factors makes in the reprogramming process and determine whether approaches that do not rely on viruses and transcription factors can have similar success.
In addition, studies are needed to test whether these results can be achieved using human cells and if other, non-blood cells can be reprogrammed to iHSCs. ![]()

Researchers have found a way to reprogram mature blood cells from mice into hematopoietic stem cells (HSCs), according to a paper published in Cell.
The team used 8 transcription factors to reprogram blood progenitor cells and mature mouse myeloid cells into HSCs.
These cells, called induced HSCs (iHSCs), have the functional hallmarks of natural HSCs, are able to self-renew like natural HSCs, and can give rise to all of the cellular components of the blood.
“Blood cell production invariably goes in one direction—from stem cells, to progenitors, to mature effector cells,” said study author Derrick J. Rossi, PhD, of Boston Children’s Hospital in Massachusetts.
“We wanted to reverse the process and derive HSCs from differentiated blood cells using transcription factors that we found were specific to HSCs.”
To that end, Dr Rossi and his colleagues screened gene expression in 40 different types of blood and blood progenitor cells from mice. From this screen, the team identified 36 transcription factors that are expressed in HSCs but not in the cells that arise from them.
In a series of mouse transplantation experiments, the researchers found that 6 of the 36 transcription factors—Hlf, Runx1t1, Pbx1, Lmo2, Zfp37, and Prdm5—plus 2 additional factors not originally identified in their screen—Mycn and Meis1—were sufficient to reprogram 2 kinds of blood progenitor cells—pro/pre-B cells and common myeloid progenitor cells—into iHSCs.
The team reprogrammed their source cells by exposing them to viruses containing the genes for all 8 transcription factors and a molecular switch that turned the factor genes on in the presence of doxycycline. They then transplanted the exposed cells into recipient mice and activated the genes by giving the mice doxycycline.
The resulting iHSCs were capable of generating the entire blood cell repertoire in the transplanted mice, showing they had gained the ability to differentiate into all blood lineages. Stem cells collected from those recipients were capable of reconstituting the blood of secondary transplant recipients, proving that the 8-factor cocktail could instill the capacity for self-renewal.
Taking the work a step further, the researchers treated mature mouse myeloid cells with the same 8-factor cocktail. The resulting iHSCs produced all of the blood lineages and could regenerate the blood of secondary transplant recipients.
Study author Stuart Orkin, MD, of the Dana-Farber Cancer Institute in Boston, noted that the use of mice as a kind of reactor for reprogramming marks a novel direction in HSC research.
“In the blood research field, no one has the conditions to expand HSCs in the tissue culture dish,” he said. “Instead, by letting the reprogramming occur in mice, Rossi takes advantage of the signaling and environmental cues HSCs would normally experience.”
Dr Orkin added that iHSCs are nearly indistinguishable from normal HSCs at the transcriptional level. Unfortunately, though, these findings are far from translation to the clinic.
Researchers must still ascertain the precise contribution each of the 8 transcription factors makes in the reprogramming process and determine whether approaches that do not rely on viruses and transcription factors can have similar success.
In addition, studies are needed to test whether these results can be achieved using human cells and if other, non-blood cells can be reprogrammed to iHSCs. ![]()

Researchers have found a way to reprogram mature blood cells from mice into hematopoietic stem cells (HSCs), according to a paper published in Cell.
The team used 8 transcription factors to reprogram blood progenitor cells and mature mouse myeloid cells into HSCs.
These cells, called induced HSCs (iHSCs), have the functional hallmarks of natural HSCs, are able to self-renew like natural HSCs, and can give rise to all of the cellular components of the blood.
“Blood cell production invariably goes in one direction—from stem cells, to progenitors, to mature effector cells,” said study author Derrick J. Rossi, PhD, of Boston Children’s Hospital in Massachusetts.
“We wanted to reverse the process and derive HSCs from differentiated blood cells using transcription factors that we found were specific to HSCs.”
To that end, Dr Rossi and his colleagues screened gene expression in 40 different types of blood and blood progenitor cells from mice. From this screen, the team identified 36 transcription factors that are expressed in HSCs but not in the cells that arise from them.
In a series of mouse transplantation experiments, the researchers found that 6 of the 36 transcription factors—Hlf, Runx1t1, Pbx1, Lmo2, Zfp37, and Prdm5—plus 2 additional factors not originally identified in their screen—Mycn and Meis1—were sufficient to reprogram 2 kinds of blood progenitor cells—pro/pre-B cells and common myeloid progenitor cells—into iHSCs.
The team reprogrammed their source cells by exposing them to viruses containing the genes for all 8 transcription factors and a molecular switch that turned the factor genes on in the presence of doxycycline. They then transplanted the exposed cells into recipient mice and activated the genes by giving the mice doxycycline.
The resulting iHSCs were capable of generating the entire blood cell repertoire in the transplanted mice, showing they had gained the ability to differentiate into all blood lineages. Stem cells collected from those recipients were capable of reconstituting the blood of secondary transplant recipients, proving that the 8-factor cocktail could instill the capacity for self-renewal.
Taking the work a step further, the researchers treated mature mouse myeloid cells with the same 8-factor cocktail. The resulting iHSCs produced all of the blood lineages and could regenerate the blood of secondary transplant recipients.
Study author Stuart Orkin, MD, of the Dana-Farber Cancer Institute in Boston, noted that the use of mice as a kind of reactor for reprogramming marks a novel direction in HSC research.
“In the blood research field, no one has the conditions to expand HSCs in the tissue culture dish,” he said. “Instead, by letting the reprogramming occur in mice, Rossi takes advantage of the signaling and environmental cues HSCs would normally experience.”
Dr Orkin added that iHSCs are nearly indistinguishable from normal HSCs at the transcriptional level. Unfortunately, though, these findings are far from translation to the clinic.
Researchers must still ascertain the precise contribution each of the 8 transcription factors makes in the reprogramming process and determine whether approaches that do not rely on viruses and transcription factors can have similar success.
In addition, studies are needed to test whether these results can be achieved using human cells and if other, non-blood cells can be reprogrammed to iHSCs. ![]()
A new method for measuring DNA repair
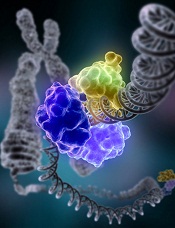
Credit: NIGMS
Cells have several major repair systems that can fix DNA damage, which may lead to cancer and other diseases if not mended.
Unfortunately, the effectiveness of these repair systems varies greatly from person to person.
Now, researchers have developed a test that can rapidly assess several of these repair systems, which could potentially help us determine an individual’s risk of developing cancer and predict how a patient might respond to chemotherapy.
The new test, described in Proceedings of the National Academy of Sciences, can analyze 4 types of DNA repair capacity simultaneously, in less than 24 hours. Previous tests have only been able to evaluate a single system at a time.
“All of the repair pathways work differently, and the existing technology to measure each of those pathways is very different for each one,” said study author Zachary Nagel, PhD, of the Massachusetts Institute of Technology in Cambridge.
“What we wanted to do was come up with one way of measuring all DNA repair pathways at the same time so you have a single readout that’s easy to measure.”
The researchers used this approach to measure DNA repair in lymphoblastoid cells taken from 24 healthy subjects. The team found a huge range of variability, especially in one repair system, where some subjects’ cells were more than 10 times more efficient than others.
“None of the cells came out looking the same,” said study author Leona Samson, PhD, also of MIT. “They each have their own spectrum of what they can repair well and what they don’t repair well. It’s like a fingerprint for each person.”
Measuring repair
With the new test, the team can measure how well cells repair the most common DNA lesions, including single-strand breaks, double-strand breaks, mismatches, and the introduction of alkyl groups caused by pollutants such as fuel exhaust and tobacco smoke.
To achieve this, the researchers created 5 different circular pieces of DNA, 4 of which carry DNA lesions. Each of these circular DNA strands, or plasmids, also carries a gene for a different colored fluorescent protein.
In some cases, the DNA lesions prevent those genes from being expressed, so when the DNA is successfully repaired, the cell begins to produce the fluorescent protein. In others, repairing the DNA lesion turns the fluorescent gene off.
By introducing these plasmids into cells and reading the fluorescent output, scientists can determine how efficiently each kind of lesion has been repaired. In theory, more than 5 plasmids could go into each cell, but the researchers limited each experiment to 5 reporter plasmids to avoid potential overlap among colors.
To overcome that limitation, the researchers are also developing an alternative tactic that involves sequencing the messenger RNA produced by cells when they copy the plasmid genes, instead of measuring fluorescence.
In this study, the team tested the sequencing approach with just one type of DNA repair, but it could allow for unlimited tests at one time. And the researchers could customize the target DNA sequence to reveal information about which type of lesion the plasmid carries, as well as information about which patient’s cells are being tested.
This would provide the ability for many different patient samples to be tested in the same batch, making the test more cost-effective.
Making predictions
Previous studies have shown that many different types of DNA repair capacity can vary greatly among apparently healthy individuals. Some of these differences have been linked with cancer vulnerability.
Scientists have also identified links between DNA repair and neurological, developmental, and immunological disorders. But useful predictive DNA-repair-based tests have not been developed, largely because it has been impossible to rapidly analyze several different types of DNA repair capacity at once.
Dr Samson’s lab is now working on adapting the new test so it can be used with blood samples taken from patients, allowing researchers to identify patients who are at higher risk of disease and potentially enabling prevention or earlier diagnosis of diseases linked to DNA repair.
Such a test could also be used to predict a patient’s response to chemotherapy or to determine how much radiation treatment a patient can tolerate.
The researchers also believe this test could be exploited to screen for new drugs that inhibit or enhance DNA repair. Inhibitors could be targeted to tumors to make them more susceptible to chemotherapy, while enhancers could help protect people who have been accidentally exposed to DNA-damaging agents, such as radiation. ![]()

Credit: NIGMS
Cells have several major repair systems that can fix DNA damage, which may lead to cancer and other diseases if not mended.
Unfortunately, the effectiveness of these repair systems varies greatly from person to person.
Now, researchers have developed a test that can rapidly assess several of these repair systems, which could potentially help us determine an individual’s risk of developing cancer and predict how a patient might respond to chemotherapy.
The new test, described in Proceedings of the National Academy of Sciences, can analyze 4 types of DNA repair capacity simultaneously, in less than 24 hours. Previous tests have only been able to evaluate a single system at a time.
“All of the repair pathways work differently, and the existing technology to measure each of those pathways is very different for each one,” said study author Zachary Nagel, PhD, of the Massachusetts Institute of Technology in Cambridge.
“What we wanted to do was come up with one way of measuring all DNA repair pathways at the same time so you have a single readout that’s easy to measure.”
The researchers used this approach to measure DNA repair in lymphoblastoid cells taken from 24 healthy subjects. The team found a huge range of variability, especially in one repair system, where some subjects’ cells were more than 10 times more efficient than others.
“None of the cells came out looking the same,” said study author Leona Samson, PhD, also of MIT. “They each have their own spectrum of what they can repair well and what they don’t repair well. It’s like a fingerprint for each person.”
Measuring repair
With the new test, the team can measure how well cells repair the most common DNA lesions, including single-strand breaks, double-strand breaks, mismatches, and the introduction of alkyl groups caused by pollutants such as fuel exhaust and tobacco smoke.
To achieve this, the researchers created 5 different circular pieces of DNA, 4 of which carry DNA lesions. Each of these circular DNA strands, or plasmids, also carries a gene for a different colored fluorescent protein.
In some cases, the DNA lesions prevent those genes from being expressed, so when the DNA is successfully repaired, the cell begins to produce the fluorescent protein. In others, repairing the DNA lesion turns the fluorescent gene off.
By introducing these plasmids into cells and reading the fluorescent output, scientists can determine how efficiently each kind of lesion has been repaired. In theory, more than 5 plasmids could go into each cell, but the researchers limited each experiment to 5 reporter plasmids to avoid potential overlap among colors.
To overcome that limitation, the researchers are also developing an alternative tactic that involves sequencing the messenger RNA produced by cells when they copy the plasmid genes, instead of measuring fluorescence.
In this study, the team tested the sequencing approach with just one type of DNA repair, but it could allow for unlimited tests at one time. And the researchers could customize the target DNA sequence to reveal information about which type of lesion the plasmid carries, as well as information about which patient’s cells are being tested.
This would provide the ability for many different patient samples to be tested in the same batch, making the test more cost-effective.
Making predictions
Previous studies have shown that many different types of DNA repair capacity can vary greatly among apparently healthy individuals. Some of these differences have been linked with cancer vulnerability.
Scientists have also identified links between DNA repair and neurological, developmental, and immunological disorders. But useful predictive DNA-repair-based tests have not been developed, largely because it has been impossible to rapidly analyze several different types of DNA repair capacity at once.
Dr Samson’s lab is now working on adapting the new test so it can be used with blood samples taken from patients, allowing researchers to identify patients who are at higher risk of disease and potentially enabling prevention or earlier diagnosis of diseases linked to DNA repair.
Such a test could also be used to predict a patient’s response to chemotherapy or to determine how much radiation treatment a patient can tolerate.
The researchers also believe this test could be exploited to screen for new drugs that inhibit or enhance DNA repair. Inhibitors could be targeted to tumors to make them more susceptible to chemotherapy, while enhancers could help protect people who have been accidentally exposed to DNA-damaging agents, such as radiation. ![]()

Credit: NIGMS
Cells have several major repair systems that can fix DNA damage, which may lead to cancer and other diseases if not mended.
Unfortunately, the effectiveness of these repair systems varies greatly from person to person.
Now, researchers have developed a test that can rapidly assess several of these repair systems, which could potentially help us determine an individual’s risk of developing cancer and predict how a patient might respond to chemotherapy.
The new test, described in Proceedings of the National Academy of Sciences, can analyze 4 types of DNA repair capacity simultaneously, in less than 24 hours. Previous tests have only been able to evaluate a single system at a time.
“All of the repair pathways work differently, and the existing technology to measure each of those pathways is very different for each one,” said study author Zachary Nagel, PhD, of the Massachusetts Institute of Technology in Cambridge.
“What we wanted to do was come up with one way of measuring all DNA repair pathways at the same time so you have a single readout that’s easy to measure.”
The researchers used this approach to measure DNA repair in lymphoblastoid cells taken from 24 healthy subjects. The team found a huge range of variability, especially in one repair system, where some subjects’ cells were more than 10 times more efficient than others.
“None of the cells came out looking the same,” said study author Leona Samson, PhD, also of MIT. “They each have their own spectrum of what they can repair well and what they don’t repair well. It’s like a fingerprint for each person.”
Measuring repair
With the new test, the team can measure how well cells repair the most common DNA lesions, including single-strand breaks, double-strand breaks, mismatches, and the introduction of alkyl groups caused by pollutants such as fuel exhaust and tobacco smoke.
To achieve this, the researchers created 5 different circular pieces of DNA, 4 of which carry DNA lesions. Each of these circular DNA strands, or plasmids, also carries a gene for a different colored fluorescent protein.
In some cases, the DNA lesions prevent those genes from being expressed, so when the DNA is successfully repaired, the cell begins to produce the fluorescent protein. In others, repairing the DNA lesion turns the fluorescent gene off.
By introducing these plasmids into cells and reading the fluorescent output, scientists can determine how efficiently each kind of lesion has been repaired. In theory, more than 5 plasmids could go into each cell, but the researchers limited each experiment to 5 reporter plasmids to avoid potential overlap among colors.
To overcome that limitation, the researchers are also developing an alternative tactic that involves sequencing the messenger RNA produced by cells when they copy the plasmid genes, instead of measuring fluorescence.
In this study, the team tested the sequencing approach with just one type of DNA repair, but it could allow for unlimited tests at one time. And the researchers could customize the target DNA sequence to reveal information about which type of lesion the plasmid carries, as well as information about which patient’s cells are being tested.
This would provide the ability for many different patient samples to be tested in the same batch, making the test more cost-effective.
Making predictions
Previous studies have shown that many different types of DNA repair capacity can vary greatly among apparently healthy individuals. Some of these differences have been linked with cancer vulnerability.
Scientists have also identified links between DNA repair and neurological, developmental, and immunological disorders. But useful predictive DNA-repair-based tests have not been developed, largely because it has been impossible to rapidly analyze several different types of DNA repair capacity at once.
Dr Samson’s lab is now working on adapting the new test so it can be used with blood samples taken from patients, allowing researchers to identify patients who are at higher risk of disease and potentially enabling prevention or earlier diagnosis of diseases linked to DNA repair.
Such a test could also be used to predict a patient’s response to chemotherapy or to determine how much radiation treatment a patient can tolerate.
The researchers also believe this test could be exploited to screen for new drugs that inhibit or enhance DNA repair. Inhibitors could be targeted to tumors to make them more susceptible to chemotherapy, while enhancers could help protect people who have been accidentally exposed to DNA-damaging agents, such as radiation. ![]()
Group maps B-cell development
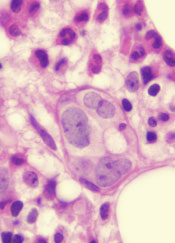
New technology has allowed scientists to create the most comprehensive map of B-cell development to date, according to a paper published in Cell.
The team combined emerging technologies for studying single cells with an advanced computational algorithm to map human B-cell development.
They believe their approach could improve researchers’ ability to investigate development in all cells and make it possible to identify rare aberrations that lead to disease.
“There are so many diseases that result from malfunctions in the molecular programs that control the development of our cell repertoire and so many rare, yet important, regulatory cell types that we have yet to discover,” said study author Dana Pe’er, PhD, of Columbia University in New York.
“We can only truly understand what goes wrong in these diseases if we have a complete map of the progression in normal development.”
Combining technologies
Dr Pe’er and her colleagues used mass cytology to observe cells in a bone marrow sample. In a single experiment, mass cytology can measure 44 molecular markers simultaneously in millions of individual cells. This provides data that can be used to compare, categorize, and order cells, as well as identify the molecular systems responsible for development.
Taking advantage of this data required the researchers to develop new mathematical and computational methods for interpreting it. Just as one can represent a physical object in 3 dimensions, the Pe’er lab’s approach involved thinking of the 44 measurements as a 44-dimensional geometric object.
So they created a new computational algorithm called Wanderlust, which uses mathematical concepts from a field called graph theory to reduce this high-dimensional data into a simple form that is easier to interpret. Wanderlust converts the developmental marker measurements in each cell into a single, 1-dimensional value that corresponds to the cell’s place within the chronology of development.
“Our body has trillions of cells of countless different types, each type bearing different molecular features and behavior,” Dr Pe’er noted. “This complexity expands from a single cell in a carefully regulated process called development.”
“This regulation creates patterns and shapes in the high-dimensional data we measure. By using Wanderlust to analyze these data, we can find the pattern and trace the trajectory that cellular development follows.”
Mapping B-cell development
To test their approach, the researchers studied development in human B cells. The team used mass cytometry to profile 44 markers in a cohort of approximately 200,000 healthy immune cells that were gathered from a single bone marrow sample.
In each cell, they measured surface markers that help identify cell type, as well as markers inside the cell that can reveal what the cell is doing, including markers for signaling, the cell cycle, apoptosis, and genome rearrangement.
Using Wanderlust to analyze the high-dimensional data provided by mass cytometry, the researchers accurately ordered the entire trajectory of 200,000 cells according to their developmental chronology. Wanderlust captured and correctly ordered all of the primary molecular landmarks known to be present in human B-cell development.
The algorithm also pinpointed a number of previously unknown regulatory signaling checkpoints that are required for human B-cell development, as well as uncharacterized subtypes of B-cell progenitors that correspond to developmental stages.
The researchers identified rare, previously unknown signaling events involving STAT5 that occurred in just 7 out of 10,000 cells. The team found that disrupting these signaling events using kinase inhibitors fully stalled the development of B cells.
Identifying and characterizing the regulatory checkpoints that control and monitor cell fate can have many practical applications, the researchers said, including the development of new diagnostics and therapeutics.
Furthermore, the team’s mapping process can be applied to any type of cell. They believe their method offers the possibility of studying normal development as well as the processes responsible for any kind of developmental disease.
“This current project is a landmark, both in the study of development and in single-cell research, and has completely changed the way I think about science,” Dr Pe’er said. “A fire has been lit, and these findings are just the tip of the iceberg of what is now possible.” ![]()

New technology has allowed scientists to create the most comprehensive map of B-cell development to date, according to a paper published in Cell.
The team combined emerging technologies for studying single cells with an advanced computational algorithm to map human B-cell development.
They believe their approach could improve researchers’ ability to investigate development in all cells and make it possible to identify rare aberrations that lead to disease.
“There are so many diseases that result from malfunctions in the molecular programs that control the development of our cell repertoire and so many rare, yet important, regulatory cell types that we have yet to discover,” said study author Dana Pe’er, PhD, of Columbia University in New York.
“We can only truly understand what goes wrong in these diseases if we have a complete map of the progression in normal development.”
Combining technologies
Dr Pe’er and her colleagues used mass cytology to observe cells in a bone marrow sample. In a single experiment, mass cytology can measure 44 molecular markers simultaneously in millions of individual cells. This provides data that can be used to compare, categorize, and order cells, as well as identify the molecular systems responsible for development.
Taking advantage of this data required the researchers to develop new mathematical and computational methods for interpreting it. Just as one can represent a physical object in 3 dimensions, the Pe’er lab’s approach involved thinking of the 44 measurements as a 44-dimensional geometric object.
So they created a new computational algorithm called Wanderlust, which uses mathematical concepts from a field called graph theory to reduce this high-dimensional data into a simple form that is easier to interpret. Wanderlust converts the developmental marker measurements in each cell into a single, 1-dimensional value that corresponds to the cell’s place within the chronology of development.
“Our body has trillions of cells of countless different types, each type bearing different molecular features and behavior,” Dr Pe’er noted. “This complexity expands from a single cell in a carefully regulated process called development.”
“This regulation creates patterns and shapes in the high-dimensional data we measure. By using Wanderlust to analyze these data, we can find the pattern and trace the trajectory that cellular development follows.”
Mapping B-cell development
To test their approach, the researchers studied development in human B cells. The team used mass cytometry to profile 44 markers in a cohort of approximately 200,000 healthy immune cells that were gathered from a single bone marrow sample.
In each cell, they measured surface markers that help identify cell type, as well as markers inside the cell that can reveal what the cell is doing, including markers for signaling, the cell cycle, apoptosis, and genome rearrangement.
Using Wanderlust to analyze the high-dimensional data provided by mass cytometry, the researchers accurately ordered the entire trajectory of 200,000 cells according to their developmental chronology. Wanderlust captured and correctly ordered all of the primary molecular landmarks known to be present in human B-cell development.
The algorithm also pinpointed a number of previously unknown regulatory signaling checkpoints that are required for human B-cell development, as well as uncharacterized subtypes of B-cell progenitors that correspond to developmental stages.
The researchers identified rare, previously unknown signaling events involving STAT5 that occurred in just 7 out of 10,000 cells. The team found that disrupting these signaling events using kinase inhibitors fully stalled the development of B cells.
Identifying and characterizing the regulatory checkpoints that control and monitor cell fate can have many practical applications, the researchers said, including the development of new diagnostics and therapeutics.
Furthermore, the team’s mapping process can be applied to any type of cell. They believe their method offers the possibility of studying normal development as well as the processes responsible for any kind of developmental disease.
“This current project is a landmark, both in the study of development and in single-cell research, and has completely changed the way I think about science,” Dr Pe’er said. “A fire has been lit, and these findings are just the tip of the iceberg of what is now possible.” ![]()

New technology has allowed scientists to create the most comprehensive map of B-cell development to date, according to a paper published in Cell.
The team combined emerging technologies for studying single cells with an advanced computational algorithm to map human B-cell development.
They believe their approach could improve researchers’ ability to investigate development in all cells and make it possible to identify rare aberrations that lead to disease.
“There are so many diseases that result from malfunctions in the molecular programs that control the development of our cell repertoire and so many rare, yet important, regulatory cell types that we have yet to discover,” said study author Dana Pe’er, PhD, of Columbia University in New York.
“We can only truly understand what goes wrong in these diseases if we have a complete map of the progression in normal development.”
Combining technologies
Dr Pe’er and her colleagues used mass cytology to observe cells in a bone marrow sample. In a single experiment, mass cytology can measure 44 molecular markers simultaneously in millions of individual cells. This provides data that can be used to compare, categorize, and order cells, as well as identify the molecular systems responsible for development.
Taking advantage of this data required the researchers to develop new mathematical and computational methods for interpreting it. Just as one can represent a physical object in 3 dimensions, the Pe’er lab’s approach involved thinking of the 44 measurements as a 44-dimensional geometric object.
So they created a new computational algorithm called Wanderlust, which uses mathematical concepts from a field called graph theory to reduce this high-dimensional data into a simple form that is easier to interpret. Wanderlust converts the developmental marker measurements in each cell into a single, 1-dimensional value that corresponds to the cell’s place within the chronology of development.
“Our body has trillions of cells of countless different types, each type bearing different molecular features and behavior,” Dr Pe’er noted. “This complexity expands from a single cell in a carefully regulated process called development.”
“This regulation creates patterns and shapes in the high-dimensional data we measure. By using Wanderlust to analyze these data, we can find the pattern and trace the trajectory that cellular development follows.”
Mapping B-cell development
To test their approach, the researchers studied development in human B cells. The team used mass cytometry to profile 44 markers in a cohort of approximately 200,000 healthy immune cells that were gathered from a single bone marrow sample.
In each cell, they measured surface markers that help identify cell type, as well as markers inside the cell that can reveal what the cell is doing, including markers for signaling, the cell cycle, apoptosis, and genome rearrangement.
Using Wanderlust to analyze the high-dimensional data provided by mass cytometry, the researchers accurately ordered the entire trajectory of 200,000 cells according to their developmental chronology. Wanderlust captured and correctly ordered all of the primary molecular landmarks known to be present in human B-cell development.
The algorithm also pinpointed a number of previously unknown regulatory signaling checkpoints that are required for human B-cell development, as well as uncharacterized subtypes of B-cell progenitors that correspond to developmental stages.
The researchers identified rare, previously unknown signaling events involving STAT5 that occurred in just 7 out of 10,000 cells. The team found that disrupting these signaling events using kinase inhibitors fully stalled the development of B cells.
Identifying and characterizing the regulatory checkpoints that control and monitor cell fate can have many practical applications, the researchers said, including the development of new diagnostics and therapeutics.
Furthermore, the team’s mapping process can be applied to any type of cell. They believe their method offers the possibility of studying normal development as well as the processes responsible for any kind of developmental disease.
“This current project is a landmark, both in the study of development and in single-cell research, and has completely changed the way I think about science,” Dr Pe’er said. “A fire has been lit, and these findings are just the tip of the iceberg of what is now possible.” ![]()
Potential for flawed research prompts new guidelines

Credit: Rhoda Baer
Worried about the potential for false conclusions in genomics research, a group of scientists created guidelines for distinguishing disease-causing sequence variants from potentially functional variants in the human genome.
In addition to helping researchers confirm variant-disease causality, the guidelines provide recommendations pertaining to genomic study design, databases, and disease diagnosis.
The guidelines are published in Nature.
They were born out of discussions at a 2012 workshop sponsored by the National Human Genome Research Institute.
“Several of us had noticed that studies were coming out with wrong conclusions about the relationship between a specific sequence and disease, and we were extremely concerned that this would translate into inappropriate clinical decisions,” said guideline author Chris Gunter, PhD, of Marcus Autism Center in Atlanta.
In other words, the authors were worried that, based on flawed results, physicians might order additional testing or treatments that are not truly supported by a link between a genetic variant and disease.
As an example, Dr Gunter and her colleagues cited autism research. Investigators found 4 independent variations in the gene TTN when they compared genomes between individuals with and without autism.
However, the TTN gene encodes titin, the largest known protein. So variations are simply more likely in TTN than in other genes. Without applying the proper statistical corrections, researchers may have falsely concluded that TTN was worthy of further investigation in autism studies.
But Dr Gunter believes the new guidelines could help prevent such false conclusions and, therefore, inappropriate medical decisions.
She and her colleagues proposed that researchers do 2 things before concluding that a genetic variation causes a disease. First, they should perform detailed statistical analyses. Then, they should assess evidence from all sources supporting a role for the variant in that specific disease or condition.
The authors said many DNA variants “may suggest a potentially convincing story about how the variant may influence the trait,” but few will actually have causal effects. So using evidence-based guidelines is crucial.
The authors’ guidelines also highlight priorities for research and infrastructure development, including added incentives for researchers to share genetic and clinical data.
“We believe that these guidelines will be particularly useful to scientists and clinicians in other areas who want to do human genomic studies and need a defined starting point for investigating genetic effects,” Dr Gunter said. ![]()

Credit: Rhoda Baer
Worried about the potential for false conclusions in genomics research, a group of scientists created guidelines for distinguishing disease-causing sequence variants from potentially functional variants in the human genome.
In addition to helping researchers confirm variant-disease causality, the guidelines provide recommendations pertaining to genomic study design, databases, and disease diagnosis.
The guidelines are published in Nature.
They were born out of discussions at a 2012 workshop sponsored by the National Human Genome Research Institute.
“Several of us had noticed that studies were coming out with wrong conclusions about the relationship between a specific sequence and disease, and we were extremely concerned that this would translate into inappropriate clinical decisions,” said guideline author Chris Gunter, PhD, of Marcus Autism Center in Atlanta.
In other words, the authors were worried that, based on flawed results, physicians might order additional testing or treatments that are not truly supported by a link between a genetic variant and disease.
As an example, Dr Gunter and her colleagues cited autism research. Investigators found 4 independent variations in the gene TTN when they compared genomes between individuals with and without autism.
However, the TTN gene encodes titin, the largest known protein. So variations are simply more likely in TTN than in other genes. Without applying the proper statistical corrections, researchers may have falsely concluded that TTN was worthy of further investigation in autism studies.
But Dr Gunter believes the new guidelines could help prevent such false conclusions and, therefore, inappropriate medical decisions.
She and her colleagues proposed that researchers do 2 things before concluding that a genetic variation causes a disease. First, they should perform detailed statistical analyses. Then, they should assess evidence from all sources supporting a role for the variant in that specific disease or condition.
The authors said many DNA variants “may suggest a potentially convincing story about how the variant may influence the trait,” but few will actually have causal effects. So using evidence-based guidelines is crucial.
The authors’ guidelines also highlight priorities for research and infrastructure development, including added incentives for researchers to share genetic and clinical data.
“We believe that these guidelines will be particularly useful to scientists and clinicians in other areas who want to do human genomic studies and need a defined starting point for investigating genetic effects,” Dr Gunter said. ![]()

Credit: Rhoda Baer
Worried about the potential for false conclusions in genomics research, a group of scientists created guidelines for distinguishing disease-causing sequence variants from potentially functional variants in the human genome.
In addition to helping researchers confirm variant-disease causality, the guidelines provide recommendations pertaining to genomic study design, databases, and disease diagnosis.
The guidelines are published in Nature.
They were born out of discussions at a 2012 workshop sponsored by the National Human Genome Research Institute.
“Several of us had noticed that studies were coming out with wrong conclusions about the relationship between a specific sequence and disease, and we were extremely concerned that this would translate into inappropriate clinical decisions,” said guideline author Chris Gunter, PhD, of Marcus Autism Center in Atlanta.
In other words, the authors were worried that, based on flawed results, physicians might order additional testing or treatments that are not truly supported by a link between a genetic variant and disease.
As an example, Dr Gunter and her colleagues cited autism research. Investigators found 4 independent variations in the gene TTN when they compared genomes between individuals with and without autism.
However, the TTN gene encodes titin, the largest known protein. So variations are simply more likely in TTN than in other genes. Without applying the proper statistical corrections, researchers may have falsely concluded that TTN was worthy of further investigation in autism studies.
But Dr Gunter believes the new guidelines could help prevent such false conclusions and, therefore, inappropriate medical decisions.
She and her colleagues proposed that researchers do 2 things before concluding that a genetic variation causes a disease. First, they should perform detailed statistical analyses. Then, they should assess evidence from all sources supporting a role for the variant in that specific disease or condition.
The authors said many DNA variants “may suggest a potentially convincing story about how the variant may influence the trait,” but few will actually have causal effects. So using evidence-based guidelines is crucial.
The authors’ guidelines also highlight priorities for research and infrastructure development, including added incentives for researchers to share genetic and clinical data.
“We believe that these guidelines will be particularly useful to scientists and clinicians in other areas who want to do human genomic studies and need a defined starting point for investigating genetic effects,” Dr Gunter said. ![]()
Study shows how age can affect WBCs, HSCs

in the bone marrow
Whole-genome sequencing has revealed hundreds of somatic mutations in healthy white blood cells (WBCs) from a 115-year-old woman, but the mutations appear to be harmless.
The research also suggests that most of the woman’s peripheral WBCs originated from 2 related hematopoietic stem cell (HSC) clones.
And the WBCs had telomeres that were significantly shorter than telomeres from other tissues.
Study investigators said these findings indicate that the finite lifespan of HSCs, rather than the effects of somatic mutations, may lead to hematopoietic clonal evolution at “extreme ages.”
Henne Holstege, PhD, of VU University Medical Center in Amsterdam, The Netherlands, and her colleagues detailed these findings in Genome Research.
The team conducted this study to determine if, over a long lifetime, mutations can accumulate in healthy WBCs. The WBCs were donated by a supercentenarian woman, who, at the time of her death in 2005, was the oldest person in the world.
The investigators performed deep whole-genome sequencing of the woman’s WBCs. And, based on the results, they estimated that about 450 somatic mutations accumulated in the non-repetitive genome within the healthy blood compartment, which suggests that about 600 somatic mutations accumulated in the whole genome of the HSC clone.
The mutations were not tumor-derived, and only a few were detected at minimal frequencies in other tissues. The team did not detect these mutations in the brain, which rarely undergoes cell division after birth.
In addition, the mutations appeared to be tolerated by the body. And they resided primarily in non-coding regions of the genome not previously associated with disease. This included sites that are especially mutation-prone, such as methylated cytosine DNA bases and solvent-accessible stretches of DNA.
Another key finding of this research was that about 65% of the healthy blood compartment was populated by the offspring of 2 HSC clones, and 1 of these was likely derived from the other.
The investigators said a possible explanation for this oligoclonality might be found in the extremely short telomere lengths of the woman’s WBCs. The WBC telomeres were 17 times shorter than telomeres in the brain.
“Because these blood cells had extremely short telomeres, we speculate that most hematopoietic stem cells may have died from stem cell exhaustion, reaching the upper limit of stem cell divisions,” Dr Holstege said.
So the team believes the oligoclonality they observed may be a consequence of HSCs’ finite lifespan. ![]()

in the bone marrow
Whole-genome sequencing has revealed hundreds of somatic mutations in healthy white blood cells (WBCs) from a 115-year-old woman, but the mutations appear to be harmless.
The research also suggests that most of the woman’s peripheral WBCs originated from 2 related hematopoietic stem cell (HSC) clones.
And the WBCs had telomeres that were significantly shorter than telomeres from other tissues.
Study investigators said these findings indicate that the finite lifespan of HSCs, rather than the effects of somatic mutations, may lead to hematopoietic clonal evolution at “extreme ages.”
Henne Holstege, PhD, of VU University Medical Center in Amsterdam, The Netherlands, and her colleagues detailed these findings in Genome Research.
The team conducted this study to determine if, over a long lifetime, mutations can accumulate in healthy WBCs. The WBCs were donated by a supercentenarian woman, who, at the time of her death in 2005, was the oldest person in the world.
The investigators performed deep whole-genome sequencing of the woman’s WBCs. And, based on the results, they estimated that about 450 somatic mutations accumulated in the non-repetitive genome within the healthy blood compartment, which suggests that about 600 somatic mutations accumulated in the whole genome of the HSC clone.
The mutations were not tumor-derived, and only a few were detected at minimal frequencies in other tissues. The team did not detect these mutations in the brain, which rarely undergoes cell division after birth.
In addition, the mutations appeared to be tolerated by the body. And they resided primarily in non-coding regions of the genome not previously associated with disease. This included sites that are especially mutation-prone, such as methylated cytosine DNA bases and solvent-accessible stretches of DNA.
Another key finding of this research was that about 65% of the healthy blood compartment was populated by the offspring of 2 HSC clones, and 1 of these was likely derived from the other.
The investigators said a possible explanation for this oligoclonality might be found in the extremely short telomere lengths of the woman’s WBCs. The WBC telomeres were 17 times shorter than telomeres in the brain.
“Because these blood cells had extremely short telomeres, we speculate that most hematopoietic stem cells may have died from stem cell exhaustion, reaching the upper limit of stem cell divisions,” Dr Holstege said.
So the team believes the oligoclonality they observed may be a consequence of HSCs’ finite lifespan. ![]()

in the bone marrow
Whole-genome sequencing has revealed hundreds of somatic mutations in healthy white blood cells (WBCs) from a 115-year-old woman, but the mutations appear to be harmless.
The research also suggests that most of the woman’s peripheral WBCs originated from 2 related hematopoietic stem cell (HSC) clones.
And the WBCs had telomeres that were significantly shorter than telomeres from other tissues.
Study investigators said these findings indicate that the finite lifespan of HSCs, rather than the effects of somatic mutations, may lead to hematopoietic clonal evolution at “extreme ages.”
Henne Holstege, PhD, of VU University Medical Center in Amsterdam, The Netherlands, and her colleagues detailed these findings in Genome Research.
The team conducted this study to determine if, over a long lifetime, mutations can accumulate in healthy WBCs. The WBCs were donated by a supercentenarian woman, who, at the time of her death in 2005, was the oldest person in the world.
The investigators performed deep whole-genome sequencing of the woman’s WBCs. And, based on the results, they estimated that about 450 somatic mutations accumulated in the non-repetitive genome within the healthy blood compartment, which suggests that about 600 somatic mutations accumulated in the whole genome of the HSC clone.
The mutations were not tumor-derived, and only a few were detected at minimal frequencies in other tissues. The team did not detect these mutations in the brain, which rarely undergoes cell division after birth.
In addition, the mutations appeared to be tolerated by the body. And they resided primarily in non-coding regions of the genome not previously associated with disease. This included sites that are especially mutation-prone, such as methylated cytosine DNA bases and solvent-accessible stretches of DNA.
Another key finding of this research was that about 65% of the healthy blood compartment was populated by the offspring of 2 HSC clones, and 1 of these was likely derived from the other.
The investigators said a possible explanation for this oligoclonality might be found in the extremely short telomere lengths of the woman’s WBCs. The WBC telomeres were 17 times shorter than telomeres in the brain.
“Because these blood cells had extremely short telomeres, we speculate that most hematopoietic stem cells may have died from stem cell exhaustion, reaching the upper limit of stem cell divisions,” Dr Holstege said.
So the team believes the oligoclonality they observed may be a consequence of HSCs’ finite lifespan. ![]()
FDA approves first drug for multicentric Castleman’s disease
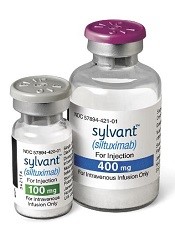
Credit: Janssen Biotech, Inc.
The US Food and Drug Administration (FDA) has authorized marketing of siltuximab (Sylvant), the first drug approved to treat patients with multicentric Castleman’s disease (MCD) who are negative for human immunodeficiency virus (HIV) and human herpes virus 8 (HHV-8).
Siltuximab is a chimeric monoclonal antibody that binds to interleukin-6 (IL-6). Dysregulated overproduction of IL-6 has been implicated in the pathogenesis of MCD.
Siltuximab has not been studied in MCD patients who are HIV- or HHV-8 positive because the drug did not bind to virally produced IL-6 in a nonclinical study.
MCD is a rare blood disorder in which lymphocytes are overproduced, leading to enlarged lymph nodes. MCD can also affect lymphoid tissue of internal organs, causing enlargement of the liver, spleen, or other organs.
Patients with MCD have a high risk of death. Infections, multisystem organ failure, and malignancies such as lymphoma are common causes of death in patients with MCD.
“There has been a serious need for treatment options for patients with MCD,” said Frits van Rhee, MD, PhD, a professor at the University of Arkansas for Medical Sciences and lead investigator of the MCD2001 study.
“MCD is a complex disease, and, up until this point, physicians have tried to reduce lymph node masses and put the disease in remission through a combination of treatments, but MCD often returns. [The approval of siltuximab] gives physicians a long-awaited treatment option for [patients] suffering with this chronic, serious, and debilitating disease.”
The FDA reviewed siltuximab under its priority review program, which provides an expedited review for drugs that demonstrate the potential to be a significant improvement in safety or effectiveness in the treatment of a serious condition. The drug was also granted orphan designation, as it is intended to treat a rare disease.
MCD2001: A phase 2 study of siltuximab
The FDA approval of siltuximab is based on results of the phase 2 MCD2001 trial. This randomized, double-blind, placebo-controlled study enrolled 79 patients with symptomatic MCD that was HIV- and HHV-8 negative.
Fifty-three patients were randomized to receive siltuximab at a dose of 11 mg/kg, plus best supportive care. The remaining 26 patients were randomized to receive placebo plus best supportive care.
The researchers defined a durable response as a tumor and symptomatic response that persisted for a minimum of 18 weeks without treatment failure. Thirty-four percent of patients in the siltuximab arm achieved this endpoint, but none of the patients in the placebo arm did (P=0.0012).
On the other hand, 4% of patients in the placebo arm experienced a tumor response, as did 38% of patients in the siltuximab arm (P<0.05).
The median time to treatment failure was not reached for patients in the siltuximab arm. But patients in the placebo arm experienced treatment failure at a median of 134 days (P<0.05).
The most frequent adverse events in siltuximab-treated patients (greater than 10% compared to placebo) were rash (28%), pruritus (28%), upper respiratory tract infection (26%), weight gain (19%), and hyperuricemia (11%).
Results of this study were presented at the 2013 ASH Annual Meeting and published in Blood. The study was sponsored by Janssen Research & Development, the company developing siltuximab.
Access to siltuximab
Siltuximab is marketed as Sylvant by Janssen Biotech Inc., which is based in Horsham, Pennsylvania. To promote access to the drug, Janssen has created the SylvantOne™ Support program.
The program offers services for providers and patients that can help assess insurance coverage and identify cost support options, such as the SylvantOne™ Patient Rebate Program for eligible commercial patients, as well as a potential option for those who are uninsured.
Patients and providers can contact SylvantOne™ Support by calling 1-855-299-8844.
Siltuximab is available in a 100-mg, single-use vial of lyophilized powder and a 400-mg, single-use vial of lyophilized powder. The recommended dose of siltuximab is 11 mg/kg given over 1 hour, via intravenous infusion, every 3 weeks.
For more information on siltuximab, see the full prescribing information. ![]()

Credit: Janssen Biotech, Inc.
The US Food and Drug Administration (FDA) has authorized marketing of siltuximab (Sylvant), the first drug approved to treat patients with multicentric Castleman’s disease (MCD) who are negative for human immunodeficiency virus (HIV) and human herpes virus 8 (HHV-8).
Siltuximab is a chimeric monoclonal antibody that binds to interleukin-6 (IL-6). Dysregulated overproduction of IL-6 has been implicated in the pathogenesis of MCD.
Siltuximab has not been studied in MCD patients who are HIV- or HHV-8 positive because the drug did not bind to virally produced IL-6 in a nonclinical study.
MCD is a rare blood disorder in which lymphocytes are overproduced, leading to enlarged lymph nodes. MCD can also affect lymphoid tissue of internal organs, causing enlargement of the liver, spleen, or other organs.
Patients with MCD have a high risk of death. Infections, multisystem organ failure, and malignancies such as lymphoma are common causes of death in patients with MCD.
“There has been a serious need for treatment options for patients with MCD,” said Frits van Rhee, MD, PhD, a professor at the University of Arkansas for Medical Sciences and lead investigator of the MCD2001 study.
“MCD is a complex disease, and, up until this point, physicians have tried to reduce lymph node masses and put the disease in remission through a combination of treatments, but MCD often returns. [The approval of siltuximab] gives physicians a long-awaited treatment option for [patients] suffering with this chronic, serious, and debilitating disease.”
The FDA reviewed siltuximab under its priority review program, which provides an expedited review for drugs that demonstrate the potential to be a significant improvement in safety or effectiveness in the treatment of a serious condition. The drug was also granted orphan designation, as it is intended to treat a rare disease.
MCD2001: A phase 2 study of siltuximab
The FDA approval of siltuximab is based on results of the phase 2 MCD2001 trial. This randomized, double-blind, placebo-controlled study enrolled 79 patients with symptomatic MCD that was HIV- and HHV-8 negative.
Fifty-three patients were randomized to receive siltuximab at a dose of 11 mg/kg, plus best supportive care. The remaining 26 patients were randomized to receive placebo plus best supportive care.
The researchers defined a durable response as a tumor and symptomatic response that persisted for a minimum of 18 weeks without treatment failure. Thirty-four percent of patients in the siltuximab arm achieved this endpoint, but none of the patients in the placebo arm did (P=0.0012).
On the other hand, 4% of patients in the placebo arm experienced a tumor response, as did 38% of patients in the siltuximab arm (P<0.05).
The median time to treatment failure was not reached for patients in the siltuximab arm. But patients in the placebo arm experienced treatment failure at a median of 134 days (P<0.05).
The most frequent adverse events in siltuximab-treated patients (greater than 10% compared to placebo) were rash (28%), pruritus (28%), upper respiratory tract infection (26%), weight gain (19%), and hyperuricemia (11%).
Results of this study were presented at the 2013 ASH Annual Meeting and published in Blood. The study was sponsored by Janssen Research & Development, the company developing siltuximab.
Access to siltuximab
Siltuximab is marketed as Sylvant by Janssen Biotech Inc., which is based in Horsham, Pennsylvania. To promote access to the drug, Janssen has created the SylvantOne™ Support program.
The program offers services for providers and patients that can help assess insurance coverage and identify cost support options, such as the SylvantOne™ Patient Rebate Program for eligible commercial patients, as well as a potential option for those who are uninsured.
Patients and providers can contact SylvantOne™ Support by calling 1-855-299-8844.
Siltuximab is available in a 100-mg, single-use vial of lyophilized powder and a 400-mg, single-use vial of lyophilized powder. The recommended dose of siltuximab is 11 mg/kg given over 1 hour, via intravenous infusion, every 3 weeks.
For more information on siltuximab, see the full prescribing information. ![]()

Credit: Janssen Biotech, Inc.
The US Food and Drug Administration (FDA) has authorized marketing of siltuximab (Sylvant), the first drug approved to treat patients with multicentric Castleman’s disease (MCD) who are negative for human immunodeficiency virus (HIV) and human herpes virus 8 (HHV-8).
Siltuximab is a chimeric monoclonal antibody that binds to interleukin-6 (IL-6). Dysregulated overproduction of IL-6 has been implicated in the pathogenesis of MCD.
Siltuximab has not been studied in MCD patients who are HIV- or HHV-8 positive because the drug did not bind to virally produced IL-6 in a nonclinical study.
MCD is a rare blood disorder in which lymphocytes are overproduced, leading to enlarged lymph nodes. MCD can also affect lymphoid tissue of internal organs, causing enlargement of the liver, spleen, or other organs.
Patients with MCD have a high risk of death. Infections, multisystem organ failure, and malignancies such as lymphoma are common causes of death in patients with MCD.
“There has been a serious need for treatment options for patients with MCD,” said Frits van Rhee, MD, PhD, a professor at the University of Arkansas for Medical Sciences and lead investigator of the MCD2001 study.
“MCD is a complex disease, and, up until this point, physicians have tried to reduce lymph node masses and put the disease in remission through a combination of treatments, but MCD often returns. [The approval of siltuximab] gives physicians a long-awaited treatment option for [patients] suffering with this chronic, serious, and debilitating disease.”
The FDA reviewed siltuximab under its priority review program, which provides an expedited review for drugs that demonstrate the potential to be a significant improvement in safety or effectiveness in the treatment of a serious condition. The drug was also granted orphan designation, as it is intended to treat a rare disease.
MCD2001: A phase 2 study of siltuximab
The FDA approval of siltuximab is based on results of the phase 2 MCD2001 trial. This randomized, double-blind, placebo-controlled study enrolled 79 patients with symptomatic MCD that was HIV- and HHV-8 negative.
Fifty-three patients were randomized to receive siltuximab at a dose of 11 mg/kg, plus best supportive care. The remaining 26 patients were randomized to receive placebo plus best supportive care.
The researchers defined a durable response as a tumor and symptomatic response that persisted for a minimum of 18 weeks without treatment failure. Thirty-four percent of patients in the siltuximab arm achieved this endpoint, but none of the patients in the placebo arm did (P=0.0012).
On the other hand, 4% of patients in the placebo arm experienced a tumor response, as did 38% of patients in the siltuximab arm (P<0.05).
The median time to treatment failure was not reached for patients in the siltuximab arm. But patients in the placebo arm experienced treatment failure at a median of 134 days (P<0.05).
The most frequent adverse events in siltuximab-treated patients (greater than 10% compared to placebo) were rash (28%), pruritus (28%), upper respiratory tract infection (26%), weight gain (19%), and hyperuricemia (11%).
Results of this study were presented at the 2013 ASH Annual Meeting and published in Blood. The study was sponsored by Janssen Research & Development, the company developing siltuximab.
Access to siltuximab
Siltuximab is marketed as Sylvant by Janssen Biotech Inc., which is based in Horsham, Pennsylvania. To promote access to the drug, Janssen has created the SylvantOne™ Support program.
The program offers services for providers and patients that can help assess insurance coverage and identify cost support options, such as the SylvantOne™ Patient Rebate Program for eligible commercial patients, as well as a potential option for those who are uninsured.
Patients and providers can contact SylvantOne™ Support by calling 1-855-299-8844.
Siltuximab is available in a 100-mg, single-use vial of lyophilized powder and a 400-mg, single-use vial of lyophilized powder. The recommended dose of siltuximab is 11 mg/kg given over 1 hour, via intravenous infusion, every 3 weeks.
For more information on siltuximab, see the full prescribing information. ![]()
FDA proposes new program for medical devices

device responds to stress
Credit: FDA
The US Food and Drug Administration (FDA) is proposing a new program to provide patients with earlier access to high-risk medical devices intended to treat or diagnose serious conditions whose medical needs are unmet by current technology.
The proposed program is called Expedited Access Premarket Approval Application for Unmet Medical Needs for Life Threatening or Irreversibly Debilitating Diseases or Conditions (shortened to EAP).
The goal of EAP is to reduce the time for premarket review of a device and the time associated with product development.
EAP allows for earlier and more interactive engagement with FDA staff, including the involvement of senior management and a plan for collecting the scientific and clinical data to support approval. The FDA says these features should help provide patients with earlier access to safe and effective medical devices.
To be eligible for participation in the program, a medical device must:
• Be intended to treat or diagnose a life-threatening or irreversibly-debilitating disease or condition
• Have an acceptable data development plan that has been approved by the FDA
• Represent 1 of the following:
1. No approved alternative treatment/diagnostic exists
2. A breakthrough technology that provides a clinically meaningful advantage over existing technology
3. Offers a significant, clinically meaningful advantage over existing approved alternatives
4. Availability is in the patient’s best interest.
In addition to the EAP, the FDA published a separate draft guidance that outlines the agency’s current policy on when data can be collected after product approval and what actions are available to the FDA if approval conditions, such as postmarket data collection, are not met.
Included in the guidance is advice on the use of surrogate or independent markers to support approval, similar to the data points used for accelerated approval of prescription drugs.
The FDA is seeking public comment on both documents:
Expedited Access for Premarket Approval Medical Devices Intended for Unmet Medical Need for Life Threatening or Irreversibly Debilitating Diseases or Conditions - Draft Guidance for Industry and FDA Staff
Balancing Premarket and Postmarket Data Collection for Devices Subject to Premarket Approval - Draft Guidance for Industry and FDA Staff. ![]()

device responds to stress
Credit: FDA
The US Food and Drug Administration (FDA) is proposing a new program to provide patients with earlier access to high-risk medical devices intended to treat or diagnose serious conditions whose medical needs are unmet by current technology.
The proposed program is called Expedited Access Premarket Approval Application for Unmet Medical Needs for Life Threatening or Irreversibly Debilitating Diseases or Conditions (shortened to EAP).
The goal of EAP is to reduce the time for premarket review of a device and the time associated with product development.
EAP allows for earlier and more interactive engagement with FDA staff, including the involvement of senior management and a plan for collecting the scientific and clinical data to support approval. The FDA says these features should help provide patients with earlier access to safe and effective medical devices.
To be eligible for participation in the program, a medical device must:
• Be intended to treat or diagnose a life-threatening or irreversibly-debilitating disease or condition
• Have an acceptable data development plan that has been approved by the FDA
• Represent 1 of the following:
1. No approved alternative treatment/diagnostic exists
2. A breakthrough technology that provides a clinically meaningful advantage over existing technology
3. Offers a significant, clinically meaningful advantage over existing approved alternatives
4. Availability is in the patient’s best interest.
In addition to the EAP, the FDA published a separate draft guidance that outlines the agency’s current policy on when data can be collected after product approval and what actions are available to the FDA if approval conditions, such as postmarket data collection, are not met.
Included in the guidance is advice on the use of surrogate or independent markers to support approval, similar to the data points used for accelerated approval of prescription drugs.
The FDA is seeking public comment on both documents:
Expedited Access for Premarket Approval Medical Devices Intended for Unmet Medical Need for Life Threatening or Irreversibly Debilitating Diseases or Conditions - Draft Guidance for Industry and FDA Staff
Balancing Premarket and Postmarket Data Collection for Devices Subject to Premarket Approval - Draft Guidance for Industry and FDA Staff. ![]()

device responds to stress
Credit: FDA
The US Food and Drug Administration (FDA) is proposing a new program to provide patients with earlier access to high-risk medical devices intended to treat or diagnose serious conditions whose medical needs are unmet by current technology.
The proposed program is called Expedited Access Premarket Approval Application for Unmet Medical Needs for Life Threatening or Irreversibly Debilitating Diseases or Conditions (shortened to EAP).
The goal of EAP is to reduce the time for premarket review of a device and the time associated with product development.
EAP allows for earlier and more interactive engagement with FDA staff, including the involvement of senior management and a plan for collecting the scientific and clinical data to support approval. The FDA says these features should help provide patients with earlier access to safe and effective medical devices.
To be eligible for participation in the program, a medical device must:
• Be intended to treat or diagnose a life-threatening or irreversibly-debilitating disease or condition
• Have an acceptable data development plan that has been approved by the FDA
• Represent 1 of the following:
1. No approved alternative treatment/diagnostic exists
2. A breakthrough technology that provides a clinically meaningful advantage over existing technology
3. Offers a significant, clinically meaningful advantage over existing approved alternatives
4. Availability is in the patient’s best interest.
In addition to the EAP, the FDA published a separate draft guidance that outlines the agency’s current policy on when data can be collected after product approval and what actions are available to the FDA if approval conditions, such as postmarket data collection, are not met.
Included in the guidance is advice on the use of surrogate or independent markers to support approval, similar to the data points used for accelerated approval of prescription drugs.
The FDA is seeking public comment on both documents:
Expedited Access for Premarket Approval Medical Devices Intended for Unmet Medical Need for Life Threatening or Irreversibly Debilitating Diseases or Conditions - Draft Guidance for Industry and FDA Staff
Balancing Premarket and Postmarket Data Collection for Devices Subject to Premarket Approval - Draft Guidance for Industry and FDA Staff.
Protein discovery could aid antimalarial development
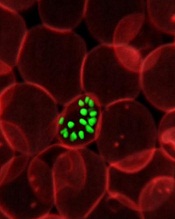
infecting a red blood cell
Credit: St Jude Hospital
High-resolution technology has revealed the structure of actin proteins in the malaria parasite Plasmodium falciparum.
Researchers found the 2 versions of the protein differ from each other more than actin proteins in any other organism.
And they were able to identify areas within these proteins that cause their different behaviors.
The team believes this discovery, published in PLOS Pathogens, may contribute to the development of tailor-made drugs against malaria.
Malaria parasites use actin to enter human cells and leave them again. Inside cells, actin confers stability, allows for cell division, and makes the movement of single cells possible.
The dynamic behavior needed for these processes is enabled by individual globular actin molecules assembling together to form thread-like structures called filaments.
The malaria parasite has 2 versions of actin, actin I and actin II, which differ substantially from each other. These structural proteins are crucial for the pathogen’s infectivity, but researchers have not been able to demonstrate filament formation in the parasite, until now.
Inari Kursula, PhD, of the University of Oulu in Finland, and her colleagues have succeeded in detecting filament assembly of the parasite actin II proteins. For this, they used electron microscopy, which overcomes the resolution limit of classical light microscopy.
Male malaria parasites from which the researchers had deleted actin II were not able to form mature germ cells and, consequently, could not reproduce and propagate.
The researchers therefore speculated that having only 1 actin variant is not sufficient for this process, but they wondered why the 2 proteins show such different behavior. To answer this question, the team deciphered the structure of the globular actin proteins using X-radiation.
“We were able to determine the structures of actin I and actin II at very high resolutions–down to 1.3 and 2.2 Ångström, respectively,” Dr Kursula said.
“With this, we are in the range of single atoms. The structures show us that the 2 variants differ more from each other than actins in any other known living organism do.”
The high resolution enabled the researchers to identify areas within the proteins that cause the different behavior.
“We now understand that Plasmodium actin filaments are very different from other actin filaments—like, for example, from those found in humans—and that they are assembled in a very different manner,” Dr Kursula said.
“Now that we know the structural basis for this, we can look for ways to specifically interfere with the parasite cytoskeleton.”
This knowledge might, in the future, aid the design of tailor-made antimalarial drugs.

infecting a red blood cell
Credit: St Jude Hospital
High-resolution technology has revealed the structure of actin proteins in the malaria parasite Plasmodium falciparum.
Researchers found the 2 versions of the protein differ from each other more than actin proteins in any other organism.
And they were able to identify areas within these proteins that cause their different behaviors.
The team believes this discovery, published in PLOS Pathogens, may contribute to the development of tailor-made drugs against malaria.
Malaria parasites use actin to enter human cells and leave them again. Inside cells, actin confers stability, allows for cell division, and makes the movement of single cells possible.
The dynamic behavior needed for these processes is enabled by individual globular actin molecules assembling together to form thread-like structures called filaments.
The malaria parasite has 2 versions of actin, actin I and actin II, which differ substantially from each other. These structural proteins are crucial for the pathogen’s infectivity, but researchers have not been able to demonstrate filament formation in the parasite, until now.
Inari Kursula, PhD, of the University of Oulu in Finland, and her colleagues have succeeded in detecting filament assembly of the parasite actin II proteins. For this, they used electron microscopy, which overcomes the resolution limit of classical light microscopy.
Male malaria parasites from which the researchers had deleted actin II were not able to form mature germ cells and, consequently, could not reproduce and propagate.
The researchers therefore speculated that having only 1 actin variant is not sufficient for this process, but they wondered why the 2 proteins show such different behavior. To answer this question, the team deciphered the structure of the globular actin proteins using X-radiation.
“We were able to determine the structures of actin I and actin II at very high resolutions–down to 1.3 and 2.2 Ångström, respectively,” Dr Kursula said.
“With this, we are in the range of single atoms. The structures show us that the 2 variants differ more from each other than actins in any other known living organism do.”
The high resolution enabled the researchers to identify areas within the proteins that cause the different behavior.
“We now understand that Plasmodium actin filaments are very different from other actin filaments—like, for example, from those found in humans—and that they are assembled in a very different manner,” Dr Kursula said.
“Now that we know the structural basis for this, we can look for ways to specifically interfere with the parasite cytoskeleton.”
This knowledge might, in the future, aid the design of tailor-made antimalarial drugs.

infecting a red blood cell
Credit: St Jude Hospital
High-resolution technology has revealed the structure of actin proteins in the malaria parasite Plasmodium falciparum.
Researchers found the 2 versions of the protein differ from each other more than actin proteins in any other organism.
And they were able to identify areas within these proteins that cause their different behaviors.
The team believes this discovery, published in PLOS Pathogens, may contribute to the development of tailor-made drugs against malaria.
Malaria parasites use actin to enter human cells and leave them again. Inside cells, actin confers stability, allows for cell division, and makes the movement of single cells possible.
The dynamic behavior needed for these processes is enabled by individual globular actin molecules assembling together to form thread-like structures called filaments.
The malaria parasite has 2 versions of actin, actin I and actin II, which differ substantially from each other. These structural proteins are crucial for the pathogen’s infectivity, but researchers have not been able to demonstrate filament formation in the parasite, until now.
Inari Kursula, PhD, of the University of Oulu in Finland, and her colleagues have succeeded in detecting filament assembly of the parasite actin II proteins. For this, they used electron microscopy, which overcomes the resolution limit of classical light microscopy.
Male malaria parasites from which the researchers had deleted actin II were not able to form mature germ cells and, consequently, could not reproduce and propagate.
The researchers therefore speculated that having only 1 actin variant is not sufficient for this process, but they wondered why the 2 proteins show such different behavior. To answer this question, the team deciphered the structure of the globular actin proteins using X-radiation.
“We were able to determine the structures of actin I and actin II at very high resolutions–down to 1.3 and 2.2 Ångström, respectively,” Dr Kursula said.
“With this, we are in the range of single atoms. The structures show us that the 2 variants differ more from each other than actins in any other known living organism do.”
The high resolution enabled the researchers to identify areas within the proteins that cause the different behavior.
“We now understand that Plasmodium actin filaments are very different from other actin filaments—like, for example, from those found in humans—and that they are assembled in a very different manner,” Dr Kursula said.
“Now that we know the structural basis for this, we can look for ways to specifically interfere with the parasite cytoskeleton.”
This knowledge might, in the future, aid the design of tailor-made antimalarial drugs.
Method speeds up analysis of RNA-seq data

Credit: Darren Baker
Researchers say they’ve developed a computational method that dramatically speeds up estimates of gene activity from RNA sequencing (RNA-seq) data.
With the new method, called Sailfish, estimates of gene expression that previously took hours can be completed in a few minutes.
And the researchers say the accuracy of Sailfish equals or exceeds that of previous methods.
The team described the Sailfish method in Nature Biotechnology.
They noted that gigantic repositories of RNA-seq data now exist, making it possible to re-analyze experiments in light of new discoveries.
“But 15 hours a pop really starts to add up, particularly if you want to look at 100 experiments,” said study author Carl Kingsford, PhD, of Carnegie Mellon University in Pittsburg, Pennsylvania.
“With Sailfish, we can give researchers everything they got from previous methods, but faster.”
With previous methods, the RNA molecules from which read sequences originated could be identified and measured only by mapping the reads to their original positions in the larger molecules—a time-consuming process.
Dr Kingsford and his colleagues found this step can actually be eliminated. The team discovered they could allocate parts of the reads to different types of RNA molecules, much as if each read acted as several “votes” for one molecule or another.
Without the mapping step, Sailfish can complete its RNA analysis 20 to 30 times faster than previous methods.
Dr Kingsford also said the Sailfish method is more robust than previous methods. It’s better able to tolerate errors in the reads or differences between individuals’ genomes.
These errors can prevent some reads from being mapped, he explained. But the Sailfish method can make use of all the RNA read “votes,” which improves the method’s accuracy.
For more information and to download the Sailfish code, visit: http://www.cs.cmu.edu/~ckingsf/software/sailfish/.

Credit: Darren Baker
Researchers say they’ve developed a computational method that dramatically speeds up estimates of gene activity from RNA sequencing (RNA-seq) data.
With the new method, called Sailfish, estimates of gene expression that previously took hours can be completed in a few minutes.
And the researchers say the accuracy of Sailfish equals or exceeds that of previous methods.
The team described the Sailfish method in Nature Biotechnology.
They noted that gigantic repositories of RNA-seq data now exist, making it possible to re-analyze experiments in light of new discoveries.
“But 15 hours a pop really starts to add up, particularly if you want to look at 100 experiments,” said study author Carl Kingsford, PhD, of Carnegie Mellon University in Pittsburg, Pennsylvania.
“With Sailfish, we can give researchers everything they got from previous methods, but faster.”
With previous methods, the RNA molecules from which read sequences originated could be identified and measured only by mapping the reads to their original positions in the larger molecules—a time-consuming process.
Dr Kingsford and his colleagues found this step can actually be eliminated. The team discovered they could allocate parts of the reads to different types of RNA molecules, much as if each read acted as several “votes” for one molecule or another.
Without the mapping step, Sailfish can complete its RNA analysis 20 to 30 times faster than previous methods.
Dr Kingsford also said the Sailfish method is more robust than previous methods. It’s better able to tolerate errors in the reads or differences between individuals’ genomes.
These errors can prevent some reads from being mapped, he explained. But the Sailfish method can make use of all the RNA read “votes,” which improves the method’s accuracy.
For more information and to download the Sailfish code, visit: http://www.cs.cmu.edu/~ckingsf/software/sailfish/.

Credit: Darren Baker
Researchers say they’ve developed a computational method that dramatically speeds up estimates of gene activity from RNA sequencing (RNA-seq) data.
With the new method, called Sailfish, estimates of gene expression that previously took hours can be completed in a few minutes.
And the researchers say the accuracy of Sailfish equals or exceeds that of previous methods.
The team described the Sailfish method in Nature Biotechnology.
They noted that gigantic repositories of RNA-seq data now exist, making it possible to re-analyze experiments in light of new discoveries.
“But 15 hours a pop really starts to add up, particularly if you want to look at 100 experiments,” said study author Carl Kingsford, PhD, of Carnegie Mellon University in Pittsburg, Pennsylvania.
“With Sailfish, we can give researchers everything they got from previous methods, but faster.”
With previous methods, the RNA molecules from which read sequences originated could be identified and measured only by mapping the reads to their original positions in the larger molecules—a time-consuming process.
Dr Kingsford and his colleagues found this step can actually be eliminated. The team discovered they could allocate parts of the reads to different types of RNA molecules, much as if each read acted as several “votes” for one molecule or another.
Without the mapping step, Sailfish can complete its RNA analysis 20 to 30 times faster than previous methods.
Dr Kingsford also said the Sailfish method is more robust than previous methods. It’s better able to tolerate errors in the reads or differences between individuals’ genomes.
These errors can prevent some reads from being mapped, he explained. But the Sailfish method can make use of all the RNA read “votes,” which improves the method’s accuracy.
For more information and to download the Sailfish code, visit: http://www.cs.cmu.edu/~ckingsf/software/sailfish/.
Study may explain how CSCs survive treatment

Credit: Andre Karwath
Experiments conducted in fruit flies showed that when researchers eliminated a type of stem cell, a group of non-stem cells stepped in to replace them.
The team said this discovery sheds new light on stem cell niches and may help explain how cancer stem cells (CSCs) replenish themselves after exposure to radiation and chemotherapy.
Erika Matunis, PhD, of the Johns Hopkins University School of Medicine in Baltimore, Maryland, and her colleagues detailed these findings in Cell Reports.
The researchers used the fruit fly as a model to examine stem cells in their natural state, studying stem cell niches in Drosophila testes.
In these niches are 3 kinds of cells: germ-line stem cells, which divide to produce sperm; somatic cyst stem cells, which make cyst cells; and hub cells, which produce signals that keep these 2 cell types going.
The hub cells have settled on their final form and are incapable of dividing further or changing their function—or so everyone thought.
In a bid to determine what happens when the somatic cyst stem cells are killed off, the researchers tried to figure out how to best do away with them. They thought the task would be straightforward, but it took many combinations of different genes working together to kill the somatic cyst cells.
“When we finally figured out a way to kill all of the somatic stem cells, we thought that the rest of the tissue would probably just empty out,” Dr Matunis said.
In 35% of testes, that’s just what happened. But in the rest, the somatic stem cells grew back.
This was a surprise, Dr Matunis said, and it raised the question of where these new stem cells originated.
The answer was another surprise: the hub cells. When the somatic stem cells were destroyed, the hub cells ramped up their machinery for cell division.
The team did several experiments to confirm the hub cells were involved, including one in which they genetically marked the hub cells and saw the mark appear in the newly formed somatic stem cells—a clear sign that hub cells had divided to make new stem cells.
Dr Matunis noted, however, that the new stem cells created by the hub cells weren’t exactly the same as the old ones. Sometimes, the new cells made molecules that only hub cells normally make.
As the researchers looked closer, they realized the damaged and recovered testes were making new niches. Instead of just one pocket of stem cells, a damaged testis might have 2 or 3.
The researchers have not determined how the new niches are formed, but they speculate that the original niche gets bigger as the new cells divide, then splits. The group is now conducting more experiments aimed at explaining the basics of how niches work, according to Dr Matunis.
She said this research may be useful for understanding CSCs. Knowing how tumor niches support the continued growth and division of CSCs might one day offer new targets for controlling such growth.

Credit: Andre Karwath
Experiments conducted in fruit flies showed that when researchers eliminated a type of stem cell, a group of non-stem cells stepped in to replace them.
The team said this discovery sheds new light on stem cell niches and may help explain how cancer stem cells (CSCs) replenish themselves after exposure to radiation and chemotherapy.
Erika Matunis, PhD, of the Johns Hopkins University School of Medicine in Baltimore, Maryland, and her colleagues detailed these findings in Cell Reports.
The researchers used the fruit fly as a model to examine stem cells in their natural state, studying stem cell niches in Drosophila testes.
In these niches are 3 kinds of cells: germ-line stem cells, which divide to produce sperm; somatic cyst stem cells, which make cyst cells; and hub cells, which produce signals that keep these 2 cell types going.
The hub cells have settled on their final form and are incapable of dividing further or changing their function—or so everyone thought.
In a bid to determine what happens when the somatic cyst stem cells are killed off, the researchers tried to figure out how to best do away with them. They thought the task would be straightforward, but it took many combinations of different genes working together to kill the somatic cyst cells.
“When we finally figured out a way to kill all of the somatic stem cells, we thought that the rest of the tissue would probably just empty out,” Dr Matunis said.
In 35% of testes, that’s just what happened. But in the rest, the somatic stem cells grew back.
This was a surprise, Dr Matunis said, and it raised the question of where these new stem cells originated.
The answer was another surprise: the hub cells. When the somatic stem cells were destroyed, the hub cells ramped up their machinery for cell division.
The team did several experiments to confirm the hub cells were involved, including one in which they genetically marked the hub cells and saw the mark appear in the newly formed somatic stem cells—a clear sign that hub cells had divided to make new stem cells.
Dr Matunis noted, however, that the new stem cells created by the hub cells weren’t exactly the same as the old ones. Sometimes, the new cells made molecules that only hub cells normally make.
As the researchers looked closer, they realized the damaged and recovered testes were making new niches. Instead of just one pocket of stem cells, a damaged testis might have 2 or 3.
The researchers have not determined how the new niches are formed, but they speculate that the original niche gets bigger as the new cells divide, then splits. The group is now conducting more experiments aimed at explaining the basics of how niches work, according to Dr Matunis.
She said this research may be useful for understanding CSCs. Knowing how tumor niches support the continued growth and division of CSCs might one day offer new targets for controlling such growth.

Credit: Andre Karwath
Experiments conducted in fruit flies showed that when researchers eliminated a type of stem cell, a group of non-stem cells stepped in to replace them.
The team said this discovery sheds new light on stem cell niches and may help explain how cancer stem cells (CSCs) replenish themselves after exposure to radiation and chemotherapy.
Erika Matunis, PhD, of the Johns Hopkins University School of Medicine in Baltimore, Maryland, and her colleagues detailed these findings in Cell Reports.
The researchers used the fruit fly as a model to examine stem cells in their natural state, studying stem cell niches in Drosophila testes.
In these niches are 3 kinds of cells: germ-line stem cells, which divide to produce sperm; somatic cyst stem cells, which make cyst cells; and hub cells, which produce signals that keep these 2 cell types going.
The hub cells have settled on their final form and are incapable of dividing further or changing their function—or so everyone thought.
In a bid to determine what happens when the somatic cyst stem cells are killed off, the researchers tried to figure out how to best do away with them. They thought the task would be straightforward, but it took many combinations of different genes working together to kill the somatic cyst cells.
“When we finally figured out a way to kill all of the somatic stem cells, we thought that the rest of the tissue would probably just empty out,” Dr Matunis said.
In 35% of testes, that’s just what happened. But in the rest, the somatic stem cells grew back.
This was a surprise, Dr Matunis said, and it raised the question of where these new stem cells originated.
The answer was another surprise: the hub cells. When the somatic stem cells were destroyed, the hub cells ramped up their machinery for cell division.
The team did several experiments to confirm the hub cells were involved, including one in which they genetically marked the hub cells and saw the mark appear in the newly formed somatic stem cells—a clear sign that hub cells had divided to make new stem cells.
Dr Matunis noted, however, that the new stem cells created by the hub cells weren’t exactly the same as the old ones. Sometimes, the new cells made molecules that only hub cells normally make.
As the researchers looked closer, they realized the damaged and recovered testes were making new niches. Instead of just one pocket of stem cells, a damaged testis might have 2 or 3.
The researchers have not determined how the new niches are formed, but they speculate that the original niche gets bigger as the new cells divide, then splits. The group is now conducting more experiments aimed at explaining the basics of how niches work, according to Dr Matunis.
She said this research may be useful for understanding CSCs. Knowing how tumor niches support the continued growth and division of CSCs might one day offer new targets for controlling such growth.