User login
Investigation reveals ‘inappropriate data handling’ but no misconduct
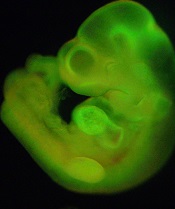
generated from STAP cells
Credit: Haruko Obokata
The Japanese research institute RIKEN has released early results of its investigation into allegations of misconduct leveled against the creators of STAP cells (stimulus-triggered acquisition of pluripotency cells).
RIKEN has confirmed 2 cases of “inappropriate data handling” but said the circumstances did not constitute misconduct.
The investigation is ongoing, with 4 issues—including charges of plagiarism and doctored figures—still to be resolved.
Research prompts questions, criticism
The investigation began shortly after a group of RIKEN scientists and colleagues from a few other institutions announced their creation of STAP cells.
The researchers said they could induce pluripotency in somatic cells by introducing the cells to a low-pH environment, and they reported this discovery in an article and a letter to Nature.
Not long after the papers were published, members of the scientific community began questioning the validity of the research, citing issues with images, possible plagiarism, and an inability to replicate the experiments described.
In light of these issues, one of the study authors recently called for the research to be retracted.
Teruhiko Wakayama, PhD, formerly of RIKEN but now a professor at the University of Yamanashi, said there are “too many uncertainties” surrounding the research at this point. After a retraction, the researchers could collect new data and images to ensure their accuracy and resubmit the research for publication.
On the other hand, fellow study author Charles Vacanti, MD, of Brigham and Women’s Hospital in Boston, has said a retraction is unnecessary.
“I firmly believe that the questions and concerns raised about our STAP cell paper published in Nature do not affect our findings or conclusions,” Dr Vacanti said.
Investigation launched
In response to the questions and allegations, RIKEN formed a committee to investigate the possibility of misconduct.
The investigation is focusing on 4 of the researchers involved: Haruko Obokata, PhD; Yoshiki Sasai, MD, PhD; Hitoshi Niwa, MD, PhD; and Dr Wakayama.
The committee is also looking into 6 issues with the research, 2 of which have been resolved.
Resolved issues
(1) Critics have questioned the “unnatural appearance of colored cell parts shown by arrows in d2 and d3 images of Figure 1f” in the article.
RIKEN concluded that the process of preparing these images did not constitute fabrication within the context of research misconduct.
(2) Questions have been raised about a “strong resemblance between the rightmost panel in Figure 1b and the lower panel in Figure 2g, both showing a fluorescence image of mice placenta” in the letter.
There is no reference to the figures in the figure legends or the main body of text, and RIKEN does define this sort of discrepancy as fabrication. However, the researchers claimed they had intended to delete one of the figures prior to publication but forgot, and there is no evidence to contradict that explanation. So RIKEN concluded that no malice was intended, and this should not be considered misconduct.
Issues under investigation
(1) In Figure 1i of the article, lane 3 appears to have been inserted.
(2) A part of the article’s “Methods” section on karyotyping analysis appears to have been copied from another paper.
(3) Some of the description of karyotyping in the “Methods” section of the article is different from the procedure the researchers followed.
(4) In the article, the image of differentiated cells for Figures 2d and 2e and the image of chimera mouse immunostaining data are incorrect, and investigation revealed that these images closely resemble images Dr Obokata used in her doctoral dissertation.
Next steps
RIKEN said it will continue with the investigation and issue a full report upon its completion. The institute also aims to determine whether the STAP cell experiments can be reproduced.
“The reproducibility and credibility of the STAP phenomenon must be rigorously validated, not only by RIKEN scientists, but also by others,” said RIKEN President Ryoji Noyori, PhD.
“I have instructed our people to cooperate fully with researchers at outside institutions in their efforts to replicate the STAP cell results.”
Dr Noyori added that RIKEN is prepared to withdraw the Nature papers and take “strict disciplinary action” against the researchers involved if the investigation reveals deliberate misconduct. ![]()

generated from STAP cells
Credit: Haruko Obokata
The Japanese research institute RIKEN has released early results of its investigation into allegations of misconduct leveled against the creators of STAP cells (stimulus-triggered acquisition of pluripotency cells).
RIKEN has confirmed 2 cases of “inappropriate data handling” but said the circumstances did not constitute misconduct.
The investigation is ongoing, with 4 issues—including charges of plagiarism and doctored figures—still to be resolved.
Research prompts questions, criticism
The investigation began shortly after a group of RIKEN scientists and colleagues from a few other institutions announced their creation of STAP cells.
The researchers said they could induce pluripotency in somatic cells by introducing the cells to a low-pH environment, and they reported this discovery in an article and a letter to Nature.
Not long after the papers were published, members of the scientific community began questioning the validity of the research, citing issues with images, possible plagiarism, and an inability to replicate the experiments described.
In light of these issues, one of the study authors recently called for the research to be retracted.
Teruhiko Wakayama, PhD, formerly of RIKEN but now a professor at the University of Yamanashi, said there are “too many uncertainties” surrounding the research at this point. After a retraction, the researchers could collect new data and images to ensure their accuracy and resubmit the research for publication.
On the other hand, fellow study author Charles Vacanti, MD, of Brigham and Women’s Hospital in Boston, has said a retraction is unnecessary.
“I firmly believe that the questions and concerns raised about our STAP cell paper published in Nature do not affect our findings or conclusions,” Dr Vacanti said.
Investigation launched
In response to the questions and allegations, RIKEN formed a committee to investigate the possibility of misconduct.
The investigation is focusing on 4 of the researchers involved: Haruko Obokata, PhD; Yoshiki Sasai, MD, PhD; Hitoshi Niwa, MD, PhD; and Dr Wakayama.
The committee is also looking into 6 issues with the research, 2 of which have been resolved.
Resolved issues
(1) Critics have questioned the “unnatural appearance of colored cell parts shown by arrows in d2 and d3 images of Figure 1f” in the article.
RIKEN concluded that the process of preparing these images did not constitute fabrication within the context of research misconduct.
(2) Questions have been raised about a “strong resemblance between the rightmost panel in Figure 1b and the lower panel in Figure 2g, both showing a fluorescence image of mice placenta” in the letter.
There is no reference to the figures in the figure legends or the main body of text, and RIKEN does define this sort of discrepancy as fabrication. However, the researchers claimed they had intended to delete one of the figures prior to publication but forgot, and there is no evidence to contradict that explanation. So RIKEN concluded that no malice was intended, and this should not be considered misconduct.
Issues under investigation
(1) In Figure 1i of the article, lane 3 appears to have been inserted.
(2) A part of the article’s “Methods” section on karyotyping analysis appears to have been copied from another paper.
(3) Some of the description of karyotyping in the “Methods” section of the article is different from the procedure the researchers followed.
(4) In the article, the image of differentiated cells for Figures 2d and 2e and the image of chimera mouse immunostaining data are incorrect, and investigation revealed that these images closely resemble images Dr Obokata used in her doctoral dissertation.
Next steps
RIKEN said it will continue with the investigation and issue a full report upon its completion. The institute also aims to determine whether the STAP cell experiments can be reproduced.
“The reproducibility and credibility of the STAP phenomenon must be rigorously validated, not only by RIKEN scientists, but also by others,” said RIKEN President Ryoji Noyori, PhD.
“I have instructed our people to cooperate fully with researchers at outside institutions in their efforts to replicate the STAP cell results.”
Dr Noyori added that RIKEN is prepared to withdraw the Nature papers and take “strict disciplinary action” against the researchers involved if the investigation reveals deliberate misconduct. ![]()

generated from STAP cells
Credit: Haruko Obokata
The Japanese research institute RIKEN has released early results of its investigation into allegations of misconduct leveled against the creators of STAP cells (stimulus-triggered acquisition of pluripotency cells).
RIKEN has confirmed 2 cases of “inappropriate data handling” but said the circumstances did not constitute misconduct.
The investigation is ongoing, with 4 issues—including charges of plagiarism and doctored figures—still to be resolved.
Research prompts questions, criticism
The investigation began shortly after a group of RIKEN scientists and colleagues from a few other institutions announced their creation of STAP cells.
The researchers said they could induce pluripotency in somatic cells by introducing the cells to a low-pH environment, and they reported this discovery in an article and a letter to Nature.
Not long after the papers were published, members of the scientific community began questioning the validity of the research, citing issues with images, possible plagiarism, and an inability to replicate the experiments described.
In light of these issues, one of the study authors recently called for the research to be retracted.
Teruhiko Wakayama, PhD, formerly of RIKEN but now a professor at the University of Yamanashi, said there are “too many uncertainties” surrounding the research at this point. After a retraction, the researchers could collect new data and images to ensure their accuracy and resubmit the research for publication.
On the other hand, fellow study author Charles Vacanti, MD, of Brigham and Women’s Hospital in Boston, has said a retraction is unnecessary.
“I firmly believe that the questions and concerns raised about our STAP cell paper published in Nature do not affect our findings or conclusions,” Dr Vacanti said.
Investigation launched
In response to the questions and allegations, RIKEN formed a committee to investigate the possibility of misconduct.
The investigation is focusing on 4 of the researchers involved: Haruko Obokata, PhD; Yoshiki Sasai, MD, PhD; Hitoshi Niwa, MD, PhD; and Dr Wakayama.
The committee is also looking into 6 issues with the research, 2 of which have been resolved.
Resolved issues
(1) Critics have questioned the “unnatural appearance of colored cell parts shown by arrows in d2 and d3 images of Figure 1f” in the article.
RIKEN concluded that the process of preparing these images did not constitute fabrication within the context of research misconduct.
(2) Questions have been raised about a “strong resemblance between the rightmost panel in Figure 1b and the lower panel in Figure 2g, both showing a fluorescence image of mice placenta” in the letter.
There is no reference to the figures in the figure legends or the main body of text, and RIKEN does define this sort of discrepancy as fabrication. However, the researchers claimed they had intended to delete one of the figures prior to publication but forgot, and there is no evidence to contradict that explanation. So RIKEN concluded that no malice was intended, and this should not be considered misconduct.
Issues under investigation
(1) In Figure 1i of the article, lane 3 appears to have been inserted.
(2) A part of the article’s “Methods” section on karyotyping analysis appears to have been copied from another paper.
(3) Some of the description of karyotyping in the “Methods” section of the article is different from the procedure the researchers followed.
(4) In the article, the image of differentiated cells for Figures 2d and 2e and the image of chimera mouse immunostaining data are incorrect, and investigation revealed that these images closely resemble images Dr Obokata used in her doctoral dissertation.
Next steps
RIKEN said it will continue with the investigation and issue a full report upon its completion. The institute also aims to determine whether the STAP cell experiments can be reproduced.
“The reproducibility and credibility of the STAP phenomenon must be rigorously validated, not only by RIKEN scientists, but also by others,” said RIKEN President Ryoji Noyori, PhD.
“I have instructed our people to cooperate fully with researchers at outside institutions in their efforts to replicate the STAP cell results.”
Dr Noyori added that RIKEN is prepared to withdraw the Nature papers and take “strict disciplinary action” against the researchers involved if the investigation reveals deliberate misconduct. ![]()
Team finds hidden reservoir of HCMV
Credit: Chad McNeeley
Researchers have found evidence suggesting that perivascular mesenchymal stromal cells (MSCs) are a reservoir of human cytomegalovirus (HCMV).
This opens up the possibility of therapeutically targeting these cells, which surround blood vessels in the organs and can be found in the bone marrow.
If effective, such a treatment method could prove life-saving for individuals who experience HCMV reactivation, such as transplant recipients and patients receiving chemotherapy.
“There are antiviral medications designed to prevent HCMV from re-activating, but HCMV infection remains one of the major complications after both organ and bone marrow transplants,” said study author Graca Almeida-Porada, MD, PhD, of Wake Forest Baptist Medical Center in Winston-Salem, North Carolina.
“The question scientists have been asking for years is, ‘Where does the virus hide when it is latent?’ Maybe if we knew, we could target it.”
Previous research showed that hematopoietic stem cells can harbor HCMV. Dr Almeida-Porada and her colleagues hypothesized that other cell populations may also harbor the virus, and they suspected that perivascular MSCs were a likely culprit.
The team’s suspicions were confirmed when testing revealed that perivascular MSCs are susceptible to HCMV infection and that the virus can grow within these cells.
The researchers also compared the susceptibility of perivascular MSCs from the liver, brain, lung, and bone marrow. And they found the highest rate of HCMV infection in cells from the lung.
“This may explain why pneumonia is the primary manifestation of the HCMV infection in bone marrow transplant recipients,” Dr Almeida-Porada said.
To expand upon these findings, she and her colleagues analyzed bone marrow samples from 19 healthy individuals who had tested positive for HCMV. Quantitative PCR revealed HCMV DNA in perivascular MSCs from 7 of the subjects.
This suggests bone marrow-derived perivascular MSCs may be a natural HCMV reservoir, according to the researchers.
“We have found another source of cells that can harbor HCMV virus,” Dr Almeida-Porada concluded. “Knowing the identity of the cells opens the possibility of targeting treatments to stop its re-activation.”
Dr Almeida-Porada and her colleagues recounted their discoveries in the American Journal of Transplantation. ![]()

Credit: Chad McNeeley
Researchers have found evidence suggesting that perivascular mesenchymal stromal cells (MSCs) are a reservoir of human cytomegalovirus (HCMV).
This opens up the possibility of therapeutically targeting these cells, which surround blood vessels in the organs and can be found in the bone marrow.
If effective, such a treatment method could prove life-saving for individuals who experience HCMV reactivation, such as transplant recipients and patients receiving chemotherapy.
“There are antiviral medications designed to prevent HCMV from re-activating, but HCMV infection remains one of the major complications after both organ and bone marrow transplants,” said study author Graca Almeida-Porada, MD, PhD, of Wake Forest Baptist Medical Center in Winston-Salem, North Carolina.
“The question scientists have been asking for years is, ‘Where does the virus hide when it is latent?’ Maybe if we knew, we could target it.”
Previous research showed that hematopoietic stem cells can harbor HCMV. Dr Almeida-Porada and her colleagues hypothesized that other cell populations may also harbor the virus, and they suspected that perivascular MSCs were a likely culprit.
The team’s suspicions were confirmed when testing revealed that perivascular MSCs are susceptible to HCMV infection and that the virus can grow within these cells.
The researchers also compared the susceptibility of perivascular MSCs from the liver, brain, lung, and bone marrow. And they found the highest rate of HCMV infection in cells from the lung.
“This may explain why pneumonia is the primary manifestation of the HCMV infection in bone marrow transplant recipients,” Dr Almeida-Porada said.
To expand upon these findings, she and her colleagues analyzed bone marrow samples from 19 healthy individuals who had tested positive for HCMV. Quantitative PCR revealed HCMV DNA in perivascular MSCs from 7 of the subjects.
This suggests bone marrow-derived perivascular MSCs may be a natural HCMV reservoir, according to the researchers.
“We have found another source of cells that can harbor HCMV virus,” Dr Almeida-Porada concluded. “Knowing the identity of the cells opens the possibility of targeting treatments to stop its re-activation.”
Dr Almeida-Porada and her colleagues recounted their discoveries in the American Journal of Transplantation. ![]()

Credit: Chad McNeeley
Researchers have found evidence suggesting that perivascular mesenchymal stromal cells (MSCs) are a reservoir of human cytomegalovirus (HCMV).
This opens up the possibility of therapeutically targeting these cells, which surround blood vessels in the organs and can be found in the bone marrow.
If effective, such a treatment method could prove life-saving for individuals who experience HCMV reactivation, such as transplant recipients and patients receiving chemotherapy.
“There are antiviral medications designed to prevent HCMV from re-activating, but HCMV infection remains one of the major complications after both organ and bone marrow transplants,” said study author Graca Almeida-Porada, MD, PhD, of Wake Forest Baptist Medical Center in Winston-Salem, North Carolina.
“The question scientists have been asking for years is, ‘Where does the virus hide when it is latent?’ Maybe if we knew, we could target it.”
Previous research showed that hematopoietic stem cells can harbor HCMV. Dr Almeida-Porada and her colleagues hypothesized that other cell populations may also harbor the virus, and they suspected that perivascular MSCs were a likely culprit.
The team’s suspicions were confirmed when testing revealed that perivascular MSCs are susceptible to HCMV infection and that the virus can grow within these cells.
The researchers also compared the susceptibility of perivascular MSCs from the liver, brain, lung, and bone marrow. And they found the highest rate of HCMV infection in cells from the lung.
“This may explain why pneumonia is the primary manifestation of the HCMV infection in bone marrow transplant recipients,” Dr Almeida-Porada said.
To expand upon these findings, she and her colleagues analyzed bone marrow samples from 19 healthy individuals who had tested positive for HCMV. Quantitative PCR revealed HCMV DNA in perivascular MSCs from 7 of the subjects.
This suggests bone marrow-derived perivascular MSCs may be a natural HCMV reservoir, according to the researchers.
“We have found another source of cells that can harbor HCMV virus,” Dr Almeida-Porada concluded. “Knowing the identity of the cells opens the possibility of targeting treatments to stop its re-activation.”
Dr Almeida-Porada and her colleagues recounted their discoveries in the American Journal of Transplantation. ![]()
Mouse model allows tracking of cells, drug activity
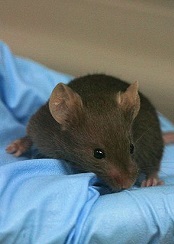
Scientists have created a mouse that expresses a fluorescing biosensor in every cell of its body, allowing diseased cells and drugs to be tracked and evaluated in real time.
The biosensor is a single-molecule probe called Raichu-Rac, which was invented in 2002.
Although Raichu-Rac has been used by many research groups since then, this is the first time researchers have successfully modified a mouse to express the molecule throughout the body without affecting cell function.
The team described this work in Cell Reports.
Raichu-Rac mimics the action of the protein Rac, which drives cell movement and oscillates between active and inactive states. When Rac is active, Raichu-Rac picks up chemical cues and glows blue. When Rac is inactive, the molecule glows yellow.
Researchers found they could use imaging techniques to follow Rac activation in any organ at any time. They could watch moment-by-moment oscillation of Rac activity at the front or back of cells as they moved in the body. And they used the technology to monitor Rac activity in response to treatment.
“It allows us to watch and map, in real time, parts of a cell or organ where Rac is active and driving invasion,” said study author Paul Timpson, PhD, of the Garvan Institute of Medical Research in Darlinghurst, New South Wales, Australia.
“In cancers, a lot of blue indicates an aggressive tumor that is in the process of spreading. You can literally watch parts of a tumor turn from blue to yellow as a drug hits its target. This can be an hour or more after the drug is administered, and the effect can wane quickly or slowly. Drug companies need to know these details—specifically, how much, how often, and how long to administer drugs.”
The researchers said the mouse can be used to study any cancer type by crossing it with other models, limiting expression of Raichu-Rac to specific cell or tissue types. And the mouse can be adapted to study other diseases by expressing Raichu-Rac in different disease models.
“The great thing about this mouse is its flexibility and potential for looking at a broad range of diseases and molecular targets,” Dr Timpson said.
The mouse was created by Heidi Welch, PhD, of the Babraham Institute in Cambridge, UK, although she said she cannot take all the credit.
“The credit must go to Professor Miki Matsuda, the genius who invented [Raichu-Rac] in the first place 12 years ago,” Dr Welch said. “He made his discovery freely available to the scientific community and has been very open about his findings since.”
“Miki Matsuda was super helpful in suggesting the expression levels we should be looking for and in recommending the exact biosensor we should use, out of many he developed.” ![]()

Scientists have created a mouse that expresses a fluorescing biosensor in every cell of its body, allowing diseased cells and drugs to be tracked and evaluated in real time.
The biosensor is a single-molecule probe called Raichu-Rac, which was invented in 2002.
Although Raichu-Rac has been used by many research groups since then, this is the first time researchers have successfully modified a mouse to express the molecule throughout the body without affecting cell function.
The team described this work in Cell Reports.
Raichu-Rac mimics the action of the protein Rac, which drives cell movement and oscillates between active and inactive states. When Rac is active, Raichu-Rac picks up chemical cues and glows blue. When Rac is inactive, the molecule glows yellow.
Researchers found they could use imaging techniques to follow Rac activation in any organ at any time. They could watch moment-by-moment oscillation of Rac activity at the front or back of cells as they moved in the body. And they used the technology to monitor Rac activity in response to treatment.
“It allows us to watch and map, in real time, parts of a cell or organ where Rac is active and driving invasion,” said study author Paul Timpson, PhD, of the Garvan Institute of Medical Research in Darlinghurst, New South Wales, Australia.
“In cancers, a lot of blue indicates an aggressive tumor that is in the process of spreading. You can literally watch parts of a tumor turn from blue to yellow as a drug hits its target. This can be an hour or more after the drug is administered, and the effect can wane quickly or slowly. Drug companies need to know these details—specifically, how much, how often, and how long to administer drugs.”
The researchers said the mouse can be used to study any cancer type by crossing it with other models, limiting expression of Raichu-Rac to specific cell or tissue types. And the mouse can be adapted to study other diseases by expressing Raichu-Rac in different disease models.
“The great thing about this mouse is its flexibility and potential for looking at a broad range of diseases and molecular targets,” Dr Timpson said.
The mouse was created by Heidi Welch, PhD, of the Babraham Institute in Cambridge, UK, although she said she cannot take all the credit.
“The credit must go to Professor Miki Matsuda, the genius who invented [Raichu-Rac] in the first place 12 years ago,” Dr Welch said. “He made his discovery freely available to the scientific community and has been very open about his findings since.”
“Miki Matsuda was super helpful in suggesting the expression levels we should be looking for and in recommending the exact biosensor we should use, out of many he developed.” ![]()

Scientists have created a mouse that expresses a fluorescing biosensor in every cell of its body, allowing diseased cells and drugs to be tracked and evaluated in real time.
The biosensor is a single-molecule probe called Raichu-Rac, which was invented in 2002.
Although Raichu-Rac has been used by many research groups since then, this is the first time researchers have successfully modified a mouse to express the molecule throughout the body without affecting cell function.
The team described this work in Cell Reports.
Raichu-Rac mimics the action of the protein Rac, which drives cell movement and oscillates between active and inactive states. When Rac is active, Raichu-Rac picks up chemical cues and glows blue. When Rac is inactive, the molecule glows yellow.
Researchers found they could use imaging techniques to follow Rac activation in any organ at any time. They could watch moment-by-moment oscillation of Rac activity at the front or back of cells as they moved in the body. And they used the technology to monitor Rac activity in response to treatment.
“It allows us to watch and map, in real time, parts of a cell or organ where Rac is active and driving invasion,” said study author Paul Timpson, PhD, of the Garvan Institute of Medical Research in Darlinghurst, New South Wales, Australia.
“In cancers, a lot of blue indicates an aggressive tumor that is in the process of spreading. You can literally watch parts of a tumor turn from blue to yellow as a drug hits its target. This can be an hour or more after the drug is administered, and the effect can wane quickly or slowly. Drug companies need to know these details—specifically, how much, how often, and how long to administer drugs.”
The researchers said the mouse can be used to study any cancer type by crossing it with other models, limiting expression of Raichu-Rac to specific cell or tissue types. And the mouse can be adapted to study other diseases by expressing Raichu-Rac in different disease models.
“The great thing about this mouse is its flexibility and potential for looking at a broad range of diseases and molecular targets,” Dr Timpson said.
The mouse was created by Heidi Welch, PhD, of the Babraham Institute in Cambridge, UK, although she said she cannot take all the credit.
“The credit must go to Professor Miki Matsuda, the genius who invented [Raichu-Rac] in the first place 12 years ago,” Dr Welch said. “He made his discovery freely available to the scientific community and has been very open about his findings since.”
“Miki Matsuda was super helpful in suggesting the expression levels we should be looking for and in recommending the exact biosensor we should use, out of many he developed.” ![]()
The challenges of whole-genome sequencing

of General Medical Sciences
There are significant challenges to overcome before whole-genome sequencing can be routinely used in the clinic, researchers have reported in JAMA.
In sequencing the entire genome of 12 healthy individuals, the team found they sometimes had difficulty identifying
variants in disease-associated genes, disagreed about which variants were particularly important for disease, and were uncertain about clinically reportable findings.
The research also revealed that detailed genome analysis required up to 100 hours of manual assessment per person, and the total cost of this process (including follow-up tests) could exceed $15,000 per person.
This is more expensive and labor-intensive than previous estimates, the researchers noted.
“We need to be very honest about what we can and cannot do at this point in time,” said study author Euan Ashley, MB ChB, DPhil, of Stanford University in California.
“Our hope is that the identification of specific hurdles will allow researchers in this field to focus their efforts on overcoming them to make this technique clinically useful.”
To identify these hurdles, Dr Ashley and his colleagues evaluated whole-genome sequencing in 12 healthy individuals. The team took note of the degree of sequencing accuracy necessary to make clinical decisions, the time it took to manually analyze each person’s results, and the costs involved.
The researchers discovered that sequencing could not provide reliable results regarding all inherited disease genes or consistently detect variants with the highest potential clinical effects.
Depending on the sequencing platform used, 10% to 19% of inherited disease genes were not covered to accepted standards for single-nucleotide variant discovery.
And although genotype concordance was high for previously described single-nucleotide variants—99% to 100%—it was low for small insertions and deletions—53% to 59%.
“It’s ironic, and slightly sobering, that we struggle the most in identifying insertions and deletions—those changes in the genome that are most impactful,” Dr Ashley said.
Nevertheless, he and his colleagues were able to identify an average of about 100 variants per person that were considered important enough for follow-up. Each variant required about an hour of investigation to assess the relevant scientific literature and determine whether the change was likely to modify disease risk in the individual.
After this process, the researchers were left with approximately 2 to 6 results they felt could be clinically important.
The variants expected to have potential health consequences were shared with 3 primary care physicians and 2 medical geneticists, who independently studied the results and recommended possible follow-up tests for each person.
The researchers estimated that the median cost for sequencing and variant interpretation was $14,815 (range, $14,050 to $15,715), plus the costs of computing infrastructure and data storage. And the projected costs for follow-up tests ranged from $351 to $776.
Despite the challenges they identified, the team said they are confident that whole-genome sequencing is worth pursuing.
“Our intention in doing this analysis was to draw a line describing where we are with this technology at this point in time and identify how best to move forward,” Dr Ashley said. “Things are becoming more clear, and the challenges to bringing this technique to the clinic are becoming crystallized.” ![]()

of General Medical Sciences
There are significant challenges to overcome before whole-genome sequencing can be routinely used in the clinic, researchers have reported in JAMA.
In sequencing the entire genome of 12 healthy individuals, the team found they sometimes had difficulty identifying
variants in disease-associated genes, disagreed about which variants were particularly important for disease, and were uncertain about clinically reportable findings.
The research also revealed that detailed genome analysis required up to 100 hours of manual assessment per person, and the total cost of this process (including follow-up tests) could exceed $15,000 per person.
This is more expensive and labor-intensive than previous estimates, the researchers noted.
“We need to be very honest about what we can and cannot do at this point in time,” said study author Euan Ashley, MB ChB, DPhil, of Stanford University in California.
“Our hope is that the identification of specific hurdles will allow researchers in this field to focus their efforts on overcoming them to make this technique clinically useful.”
To identify these hurdles, Dr Ashley and his colleagues evaluated whole-genome sequencing in 12 healthy individuals. The team took note of the degree of sequencing accuracy necessary to make clinical decisions, the time it took to manually analyze each person’s results, and the costs involved.
The researchers discovered that sequencing could not provide reliable results regarding all inherited disease genes or consistently detect variants with the highest potential clinical effects.
Depending on the sequencing platform used, 10% to 19% of inherited disease genes were not covered to accepted standards for single-nucleotide variant discovery.
And although genotype concordance was high for previously described single-nucleotide variants—99% to 100%—it was low for small insertions and deletions—53% to 59%.
“It’s ironic, and slightly sobering, that we struggle the most in identifying insertions and deletions—those changes in the genome that are most impactful,” Dr Ashley said.
Nevertheless, he and his colleagues were able to identify an average of about 100 variants per person that were considered important enough for follow-up. Each variant required about an hour of investigation to assess the relevant scientific literature and determine whether the change was likely to modify disease risk in the individual.
After this process, the researchers were left with approximately 2 to 6 results they felt could be clinically important.
The variants expected to have potential health consequences were shared with 3 primary care physicians and 2 medical geneticists, who independently studied the results and recommended possible follow-up tests for each person.
The researchers estimated that the median cost for sequencing and variant interpretation was $14,815 (range, $14,050 to $15,715), plus the costs of computing infrastructure and data storage. And the projected costs for follow-up tests ranged from $351 to $776.
Despite the challenges they identified, the team said they are confident that whole-genome sequencing is worth pursuing.
“Our intention in doing this analysis was to draw a line describing where we are with this technology at this point in time and identify how best to move forward,” Dr Ashley said. “Things are becoming more clear, and the challenges to bringing this technique to the clinic are becoming crystallized.” ![]()

of General Medical Sciences
There are significant challenges to overcome before whole-genome sequencing can be routinely used in the clinic, researchers have reported in JAMA.
In sequencing the entire genome of 12 healthy individuals, the team found they sometimes had difficulty identifying
variants in disease-associated genes, disagreed about which variants were particularly important for disease, and were uncertain about clinically reportable findings.
The research also revealed that detailed genome analysis required up to 100 hours of manual assessment per person, and the total cost of this process (including follow-up tests) could exceed $15,000 per person.
This is more expensive and labor-intensive than previous estimates, the researchers noted.
“We need to be very honest about what we can and cannot do at this point in time,” said study author Euan Ashley, MB ChB, DPhil, of Stanford University in California.
“Our hope is that the identification of specific hurdles will allow researchers in this field to focus their efforts on overcoming them to make this technique clinically useful.”
To identify these hurdles, Dr Ashley and his colleagues evaluated whole-genome sequencing in 12 healthy individuals. The team took note of the degree of sequencing accuracy necessary to make clinical decisions, the time it took to manually analyze each person’s results, and the costs involved.
The researchers discovered that sequencing could not provide reliable results regarding all inherited disease genes or consistently detect variants with the highest potential clinical effects.
Depending on the sequencing platform used, 10% to 19% of inherited disease genes were not covered to accepted standards for single-nucleotide variant discovery.
And although genotype concordance was high for previously described single-nucleotide variants—99% to 100%—it was low for small insertions and deletions—53% to 59%.
“It’s ironic, and slightly sobering, that we struggle the most in identifying insertions and deletions—those changes in the genome that are most impactful,” Dr Ashley said.
Nevertheless, he and his colleagues were able to identify an average of about 100 variants per person that were considered important enough for follow-up. Each variant required about an hour of investigation to assess the relevant scientific literature and determine whether the change was likely to modify disease risk in the individual.
After this process, the researchers were left with approximately 2 to 6 results they felt could be clinically important.
The variants expected to have potential health consequences were shared with 3 primary care physicians and 2 medical geneticists, who independently studied the results and recommended possible follow-up tests for each person.
The researchers estimated that the median cost for sequencing and variant interpretation was $14,815 (range, $14,050 to $15,715), plus the costs of computing infrastructure and data storage. And the projected costs for follow-up tests ranged from $351 to $776.
Despite the challenges they identified, the team said they are confident that whole-genome sequencing is worth pursuing.
“Our intention in doing this analysis was to draw a line describing where we are with this technology at this point in time and identify how best to move forward,” Dr Ashley said. “Things are becoming more clear, and the challenges to bringing this technique to the clinic are becoming crystallized.” ![]()
Many discontinued RCTs go unpublished, unreported

for use in a clinical trial
Credit: Esther Dyson
Too many discontinued clinical trials go unreported to ethics committees and remain unpublished, according to a paper published in JAMA.
Investigators evaluated more than 1000 randomized clinical trials (RCTs) initiated in 3 countries between 2000 and 2003. And they found that roughly a quarter of these trials were ultimately discontinued.
Although all of these RCTs had been approved by research ethics committees, less than 40% of the discontinuations were reported to the committees.
And discontinued trials were significantly more likely than completed trials to remain unpublished.
The most common reason for discontinuation was insufficient subject enrollment.
Benjamin Kasenda, MD, of University Hospital Basel in Switzerland, and his colleagues conducted this research. They examined the characteristics of 1017 RCTs approved by 6 research ethics committees in Switzerland, Germany, and Canada between 2000 and 2003. The last follow-up of these trials was April 27, 2013.
Overall, 253 RCTs (24.9%) were discontinued, but only 38% of these discontinuations were reported to ethics committees.
Discontinued trials were more likely than completed trials to remain unpublished—55.1% vs 33.6% (P<0.001).
The most common reasons for discontinuation were poor recruitment (9.9%, 101/1017), administrative reasons (3.8%, 39/1017), and futility (3.6%, 37/1017).
A multivariate analysis revealed that industry sponsorship, compared to investigator sponsorship, and a larger planned sample size in increments of 100 were associated with lower rates of discontinuation due to poor recruitment. The adjusted odds ratios were 0.25 (P<0.001) and 0.96 (P=0.04), respectively.
Dr Kasenda and his colleagues said these results suggest researchers must put forth more effort to ensure that trial discontinuation is reported to ethics committees and that results of discontinued trials are published.
They said failure to publish these results is “a waste of valid data that could contribute to systematic reviews and meta-analyses.” In addition, taking steps to improve subject recruitment could greatly reduce RCT discontinuation. ![]()

for use in a clinical trial
Credit: Esther Dyson
Too many discontinued clinical trials go unreported to ethics committees and remain unpublished, according to a paper published in JAMA.
Investigators evaluated more than 1000 randomized clinical trials (RCTs) initiated in 3 countries between 2000 and 2003. And they found that roughly a quarter of these trials were ultimately discontinued.
Although all of these RCTs had been approved by research ethics committees, less than 40% of the discontinuations were reported to the committees.
And discontinued trials were significantly more likely than completed trials to remain unpublished.
The most common reason for discontinuation was insufficient subject enrollment.
Benjamin Kasenda, MD, of University Hospital Basel in Switzerland, and his colleagues conducted this research. They examined the characteristics of 1017 RCTs approved by 6 research ethics committees in Switzerland, Germany, and Canada between 2000 and 2003. The last follow-up of these trials was April 27, 2013.
Overall, 253 RCTs (24.9%) were discontinued, but only 38% of these discontinuations were reported to ethics committees.
Discontinued trials were more likely than completed trials to remain unpublished—55.1% vs 33.6% (P<0.001).
The most common reasons for discontinuation were poor recruitment (9.9%, 101/1017), administrative reasons (3.8%, 39/1017), and futility (3.6%, 37/1017).
A multivariate analysis revealed that industry sponsorship, compared to investigator sponsorship, and a larger planned sample size in increments of 100 were associated with lower rates of discontinuation due to poor recruitment. The adjusted odds ratios were 0.25 (P<0.001) and 0.96 (P=0.04), respectively.
Dr Kasenda and his colleagues said these results suggest researchers must put forth more effort to ensure that trial discontinuation is reported to ethics committees and that results of discontinued trials are published.
They said failure to publish these results is “a waste of valid data that could contribute to systematic reviews and meta-analyses.” In addition, taking steps to improve subject recruitment could greatly reduce RCT discontinuation. ![]()

for use in a clinical trial
Credit: Esther Dyson
Too many discontinued clinical trials go unreported to ethics committees and remain unpublished, according to a paper published in JAMA.
Investigators evaluated more than 1000 randomized clinical trials (RCTs) initiated in 3 countries between 2000 and 2003. And they found that roughly a quarter of these trials were ultimately discontinued.
Although all of these RCTs had been approved by research ethics committees, less than 40% of the discontinuations were reported to the committees.
And discontinued trials were significantly more likely than completed trials to remain unpublished.
The most common reason for discontinuation was insufficient subject enrollment.
Benjamin Kasenda, MD, of University Hospital Basel in Switzerland, and his colleagues conducted this research. They examined the characteristics of 1017 RCTs approved by 6 research ethics committees in Switzerland, Germany, and Canada between 2000 and 2003. The last follow-up of these trials was April 27, 2013.
Overall, 253 RCTs (24.9%) were discontinued, but only 38% of these discontinuations were reported to ethics committees.
Discontinued trials were more likely than completed trials to remain unpublished—55.1% vs 33.6% (P<0.001).
The most common reasons for discontinuation were poor recruitment (9.9%, 101/1017), administrative reasons (3.8%, 39/1017), and futility (3.6%, 37/1017).
A multivariate analysis revealed that industry sponsorship, compared to investigator sponsorship, and a larger planned sample size in increments of 100 were associated with lower rates of discontinuation due to poor recruitment. The adjusted odds ratios were 0.25 (P<0.001) and 0.96 (P=0.04), respectively.
Dr Kasenda and his colleagues said these results suggest researchers must put forth more effort to ensure that trial discontinuation is reported to ethics committees and that results of discontinued trials are published.
They said failure to publish these results is “a waste of valid data that could contribute to systematic reviews and meta-analyses.” In addition, taking steps to improve subject recruitment could greatly reduce RCT discontinuation. ![]()
Trial registry info differs from journal publication

Credit: CDC/James Gathany
New research has revealed discrepancies between information posted on the ClinicalTrials.gov website and information published in journals.
During a 1-year period, nearly all of the trials published in “high-impact” journals and listed on ClinicalTrials.gov had at least 1 discrepancy between the 2 sources.
This included differences in study group data, intervention information, and primary and secondary endpoints.
Jessica E. Becker, of the Yale University School of Medicine in New Haven, Connecticut, and her colleagues disclosed these findings in a letter to JAMA.
The researchers identified 96 trials reporting results on ClinicalTrials.gov that were also published in 19 “high-impact” journals from July 1, 2010, to June 30, 2011. The trials were most frequently published in NEJM (n=23; 24%), The Lancet (n=18; 19%), and JAMA (n=11; 12%).
Common conditions investigated in these studies included cardiovascular disease, diabetes, and hyperlipidemia (n=21; 23%); cancer (n=20; 21%); and infectious disease (n=19; 20%). Seventy-three percent of the trials (n=70) were primarily funded by industry.
Cohort, intervention, and efficacy endpoint information was reported in both the journal and on ClinicalTrials.gov for most of the trials—ranging from 93% to 100%.
For 97% of the trials (93/96), there was at least 1 difference in information between the registry and the journal article. The level of discordance between the sources was lowest for enrollment numbers—2%—and highest for completion rates—22%.
Discordance was also quite high for the trial interventions (16%). This included differences in dosage descriptions, frequencies, and duration of the intervention.
There were 132 primary efficacy endpoints described in both sources. Fifty-two percent of these endpoints could be compared between the 2 sources and had concordant results. Results for 23% (n=30) could not be compared, and 16% (n=21) were discordant.
The majority (n=15) of discordant results did not alter the interpretation of the trial. But for 6 trials, the discordance did affect interpretation.
These trials had differences in time to disease progression, rate of disease recurrence, time to resolution of a condition, progression-free survival, and results of statistical analyses.
Among the 619 secondary efficacy endpoints that were described in both sources, results for 37% (n=228) could not be compared, and 9% (n=53) were discordant. Overall, 16% of secondary efficacy endpoints were described in both sources and reported concordant results.
The researchers said this study raises questions about the accuracy of information published on ClinicalTrials.gov and in journals.
Furthermore, because the journals studied have rigorous peer review processes, the trials in this sample may represent best-case scenarios with regard to the quality of results reporting. ![]()

Credit: CDC/James Gathany
New research has revealed discrepancies between information posted on the ClinicalTrials.gov website and information published in journals.
During a 1-year period, nearly all of the trials published in “high-impact” journals and listed on ClinicalTrials.gov had at least 1 discrepancy between the 2 sources.
This included differences in study group data, intervention information, and primary and secondary endpoints.
Jessica E. Becker, of the Yale University School of Medicine in New Haven, Connecticut, and her colleagues disclosed these findings in a letter to JAMA.
The researchers identified 96 trials reporting results on ClinicalTrials.gov that were also published in 19 “high-impact” journals from July 1, 2010, to June 30, 2011. The trials were most frequently published in NEJM (n=23; 24%), The Lancet (n=18; 19%), and JAMA (n=11; 12%).
Common conditions investigated in these studies included cardiovascular disease, diabetes, and hyperlipidemia (n=21; 23%); cancer (n=20; 21%); and infectious disease (n=19; 20%). Seventy-three percent of the trials (n=70) were primarily funded by industry.
Cohort, intervention, and efficacy endpoint information was reported in both the journal and on ClinicalTrials.gov for most of the trials—ranging from 93% to 100%.
For 97% of the trials (93/96), there was at least 1 difference in information between the registry and the journal article. The level of discordance between the sources was lowest for enrollment numbers—2%—and highest for completion rates—22%.
Discordance was also quite high for the trial interventions (16%). This included differences in dosage descriptions, frequencies, and duration of the intervention.
There were 132 primary efficacy endpoints described in both sources. Fifty-two percent of these endpoints could be compared between the 2 sources and had concordant results. Results for 23% (n=30) could not be compared, and 16% (n=21) were discordant.
The majority (n=15) of discordant results did not alter the interpretation of the trial. But for 6 trials, the discordance did affect interpretation.
These trials had differences in time to disease progression, rate of disease recurrence, time to resolution of a condition, progression-free survival, and results of statistical analyses.
Among the 619 secondary efficacy endpoints that were described in both sources, results for 37% (n=228) could not be compared, and 9% (n=53) were discordant. Overall, 16% of secondary efficacy endpoints were described in both sources and reported concordant results.
The researchers said this study raises questions about the accuracy of information published on ClinicalTrials.gov and in journals.
Furthermore, because the journals studied have rigorous peer review processes, the trials in this sample may represent best-case scenarios with regard to the quality of results reporting. ![]()

Credit: CDC/James Gathany
New research has revealed discrepancies between information posted on the ClinicalTrials.gov website and information published in journals.
During a 1-year period, nearly all of the trials published in “high-impact” journals and listed on ClinicalTrials.gov had at least 1 discrepancy between the 2 sources.
This included differences in study group data, intervention information, and primary and secondary endpoints.
Jessica E. Becker, of the Yale University School of Medicine in New Haven, Connecticut, and her colleagues disclosed these findings in a letter to JAMA.
The researchers identified 96 trials reporting results on ClinicalTrials.gov that were also published in 19 “high-impact” journals from July 1, 2010, to June 30, 2011. The trials were most frequently published in NEJM (n=23; 24%), The Lancet (n=18; 19%), and JAMA (n=11; 12%).
Common conditions investigated in these studies included cardiovascular disease, diabetes, and hyperlipidemia (n=21; 23%); cancer (n=20; 21%); and infectious disease (n=19; 20%). Seventy-three percent of the trials (n=70) were primarily funded by industry.
Cohort, intervention, and efficacy endpoint information was reported in both the journal and on ClinicalTrials.gov for most of the trials—ranging from 93% to 100%.
For 97% of the trials (93/96), there was at least 1 difference in information between the registry and the journal article. The level of discordance between the sources was lowest for enrollment numbers—2%—and highest for completion rates—22%.
Discordance was also quite high for the trial interventions (16%). This included differences in dosage descriptions, frequencies, and duration of the intervention.
There were 132 primary efficacy endpoints described in both sources. Fifty-two percent of these endpoints could be compared between the 2 sources and had concordant results. Results for 23% (n=30) could not be compared, and 16% (n=21) were discordant.
The majority (n=15) of discordant results did not alter the interpretation of the trial. But for 6 trials, the discordance did affect interpretation.
These trials had differences in time to disease progression, rate of disease recurrence, time to resolution of a condition, progression-free survival, and results of statistical analyses.
Among the 619 secondary efficacy endpoints that were described in both sources, results for 37% (n=228) could not be compared, and 9% (n=53) were discordant. Overall, 16% of secondary efficacy endpoints were described in both sources and reported concordant results.
The researchers said this study raises questions about the accuracy of information published on ClinicalTrials.gov and in journals.
Furthermore, because the journals studied have rigorous peer review processes, the trials in this sample may represent best-case scenarios with regard to the quality of results reporting. ![]()
Team uses DNA methylation to quantify leukocytes
Credit: Graham Colm
Investigators have found they can use DNA methylation to simultaneously count multiple leukocyte subsets in blood samples, and they’ve reported this finding in Genome Biology.
The researchers noted that current methods of counting leukocytes in a blood sample require whole cells.
But as their newly developed method relies on DNA, it can be useful even in archived blood samples in which cells have lost their physical integrity.
“Once you understand the unique and really immutable [methylation] signature that directs the differentiation of the cell, then you can use that, and you don’t need the cell anymore,” said study author Karl Kelsey, MD, of Brown University in Providence, Rhode Island.
So the new test detects those methylation signatures in a blood sample and, with the help of algorithms, allows researchers to count how many cells of each type are in the sample.
Dr Kelsey and his colleagues found they could quantify T cells, B cells, NK cells, monocytes, eosinophils, basophils, and neutrophils.
The investigators tested their method using fresh blood samples from more than 80 donors.
The new technique proved as accurate as 3 gold-standard methods of cell quantification: manual 5-part differential, complete blood count with automated 5-part differential, and fluorescence activated cell sorting.
In further experiments, the team found their technique could detect the mixtures of immune cells associated with known diseases.
The method also proved effective with blood exposed to storage conditions, such as freezing and the addition of anticoagulants.
Moreover, the researchers showed that to distinguish among and count those various immune cell types, they only needed to measure a few dozen methylation marks in the DNA. In other words, what’s sufficient to constitute a signature can be quite short.
This method has proven feasible enough that other epidemiology research labs are already using it, Dr Kelsey said. And Brown University has applied for a patent on the technique.
Dr Kelsey said the method has the potential to be cheaper and faster than current techniques, although the investigators didn’t test that in their experiments. ![]()

Credit: Graham Colm
Investigators have found they can use DNA methylation to simultaneously count multiple leukocyte subsets in blood samples, and they’ve reported this finding in Genome Biology.
The researchers noted that current methods of counting leukocytes in a blood sample require whole cells.
But as their newly developed method relies on DNA, it can be useful even in archived blood samples in which cells have lost their physical integrity.
“Once you understand the unique and really immutable [methylation] signature that directs the differentiation of the cell, then you can use that, and you don’t need the cell anymore,” said study author Karl Kelsey, MD, of Brown University in Providence, Rhode Island.
So the new test detects those methylation signatures in a blood sample and, with the help of algorithms, allows researchers to count how many cells of each type are in the sample.
Dr Kelsey and his colleagues found they could quantify T cells, B cells, NK cells, monocytes, eosinophils, basophils, and neutrophils.
The investigators tested their method using fresh blood samples from more than 80 donors.
The new technique proved as accurate as 3 gold-standard methods of cell quantification: manual 5-part differential, complete blood count with automated 5-part differential, and fluorescence activated cell sorting.
In further experiments, the team found their technique could detect the mixtures of immune cells associated with known diseases.
The method also proved effective with blood exposed to storage conditions, such as freezing and the addition of anticoagulants.
Moreover, the researchers showed that to distinguish among and count those various immune cell types, they only needed to measure a few dozen methylation marks in the DNA. In other words, what’s sufficient to constitute a signature can be quite short.
This method has proven feasible enough that other epidemiology research labs are already using it, Dr Kelsey said. And Brown University has applied for a patent on the technique.
Dr Kelsey said the method has the potential to be cheaper and faster than current techniques, although the investigators didn’t test that in their experiments. ![]()

Credit: Graham Colm
Investigators have found they can use DNA methylation to simultaneously count multiple leukocyte subsets in blood samples, and they’ve reported this finding in Genome Biology.
The researchers noted that current methods of counting leukocytes in a blood sample require whole cells.
But as their newly developed method relies on DNA, it can be useful even in archived blood samples in which cells have lost their physical integrity.
“Once you understand the unique and really immutable [methylation] signature that directs the differentiation of the cell, then you can use that, and you don’t need the cell anymore,” said study author Karl Kelsey, MD, of Brown University in Providence, Rhode Island.
So the new test detects those methylation signatures in a blood sample and, with the help of algorithms, allows researchers to count how many cells of each type are in the sample.
Dr Kelsey and his colleagues found they could quantify T cells, B cells, NK cells, monocytes, eosinophils, basophils, and neutrophils.
The investigators tested their method using fresh blood samples from more than 80 donors.
The new technique proved as accurate as 3 gold-standard methods of cell quantification: manual 5-part differential, complete blood count with automated 5-part differential, and fluorescence activated cell sorting.
In further experiments, the team found their technique could detect the mixtures of immune cells associated with known diseases.
The method also proved effective with blood exposed to storage conditions, such as freezing and the addition of anticoagulants.
Moreover, the researchers showed that to distinguish among and count those various immune cell types, they only needed to measure a few dozen methylation marks in the DNA. In other words, what’s sufficient to constitute a signature can be quite short.
This method has proven feasible enough that other epidemiology research labs are already using it, Dr Kelsey said. And Brown University has applied for a patent on the technique.
Dr Kelsey said the method has the potential to be cheaper and faster than current techniques, although the investigators didn’t test that in their experiments.
Rate of protein synthesis affects HSC function, study suggests

in the bone marrow
Hematopoietic stem cells (HSCs) require a highly regulated rate of protein synthesis to function properly, according to research published in Nature.
Experiments showed that a ribosomal mutation decreases protein synthesis in HSCs, and deletion of a tumor suppressor gene increases protein synthesis.
But both changes result in impaired HSC function.
In mouse models, the mutation counteracted the effects of the deletion, which restored normal HSC function and delayed leukemogenesis.
“We unveiled new areas of cellular biology that no one has seen before,” said study author Sean Morrison, PhD, of the University of Texas Southwestern Medical Center in Dallas.
“This finding not only tells us something new about stem cell regulation but opens up the ability to study differences in protein synthesis between many kinds of cells in the body. We believe there is an undiscovered world of biology that allows different kinds of cells to synthesize protein at different rates and in different ways, and that those differences are important for cellular survival.”
In a previous study, researchers discovered that, by modifying the antibiotic puromycin, they could measure protein synthesis in rare cells in vivo.
Dr Morrison and his colleagues realized they could adapt this reagent to measure protein synthesis in HSCs and other cells in the hematopoietic system.
Their analyses showed that different types of blood cells produced vastly different amounts of protein per hour. And HSCs, in particular, synthesized much less protein than other hematopoietic progenitors.
“This result suggests that blood-forming stem cells require a lower rate of protein synthesis as compared to other blood-forming cells,” Dr Morrison said.
He and his colleagues then generated mice with a mutation in a component of the ribosome (Rpl24Bst/+ mice). HSCs in these mice had a 30% lower rate of protein production than controls.
The researchers observed the opposite effect when they deleted the tumor suppressor gene Pten in mouse HSCs. These mice saw a roughly 30% increase in protein production relative to controls.
However, as in the Rpl24Bst/+ mice, HSC function was noticeably impaired in these animals.
Together, these observations suggest that HSCs require a highly regulated rate of protein synthesis, such that increases or decreases in that rate impair HSC function.
“Amazingly, when the ribosomal mutant mice and the Pten mutant mice were bred together, stem cell function returned to normal, and we greatly delayed, and in some instances entirely blocked, the development of leukemia,” Dr Morrison said.
“All of this happened because protein production in stem cells was returned to normal. It was as if two wrongs made a right.”

in the bone marrow
Hematopoietic stem cells (HSCs) require a highly regulated rate of protein synthesis to function properly, according to research published in Nature.
Experiments showed that a ribosomal mutation decreases protein synthesis in HSCs, and deletion of a tumor suppressor gene increases protein synthesis.
But both changes result in impaired HSC function.
In mouse models, the mutation counteracted the effects of the deletion, which restored normal HSC function and delayed leukemogenesis.
“We unveiled new areas of cellular biology that no one has seen before,” said study author Sean Morrison, PhD, of the University of Texas Southwestern Medical Center in Dallas.
“This finding not only tells us something new about stem cell regulation but opens up the ability to study differences in protein synthesis between many kinds of cells in the body. We believe there is an undiscovered world of biology that allows different kinds of cells to synthesize protein at different rates and in different ways, and that those differences are important for cellular survival.”
In a previous study, researchers discovered that, by modifying the antibiotic puromycin, they could measure protein synthesis in rare cells in vivo.
Dr Morrison and his colleagues realized they could adapt this reagent to measure protein synthesis in HSCs and other cells in the hematopoietic system.
Their analyses showed that different types of blood cells produced vastly different amounts of protein per hour. And HSCs, in particular, synthesized much less protein than other hematopoietic progenitors.
“This result suggests that blood-forming stem cells require a lower rate of protein synthesis as compared to other blood-forming cells,” Dr Morrison said.
He and his colleagues then generated mice with a mutation in a component of the ribosome (Rpl24Bst/+ mice). HSCs in these mice had a 30% lower rate of protein production than controls.
The researchers observed the opposite effect when they deleted the tumor suppressor gene Pten in mouse HSCs. These mice saw a roughly 30% increase in protein production relative to controls.
However, as in the Rpl24Bst/+ mice, HSC function was noticeably impaired in these animals.
Together, these observations suggest that HSCs require a highly regulated rate of protein synthesis, such that increases or decreases in that rate impair HSC function.
“Amazingly, when the ribosomal mutant mice and the Pten mutant mice were bred together, stem cell function returned to normal, and we greatly delayed, and in some instances entirely blocked, the development of leukemia,” Dr Morrison said.
“All of this happened because protein production in stem cells was returned to normal. It was as if two wrongs made a right.”

in the bone marrow
Hematopoietic stem cells (HSCs) require a highly regulated rate of protein synthesis to function properly, according to research published in Nature.
Experiments showed that a ribosomal mutation decreases protein synthesis in HSCs, and deletion of a tumor suppressor gene increases protein synthesis.
But both changes result in impaired HSC function.
In mouse models, the mutation counteracted the effects of the deletion, which restored normal HSC function and delayed leukemogenesis.
“We unveiled new areas of cellular biology that no one has seen before,” said study author Sean Morrison, PhD, of the University of Texas Southwestern Medical Center in Dallas.
“This finding not only tells us something new about stem cell regulation but opens up the ability to study differences in protein synthesis between many kinds of cells in the body. We believe there is an undiscovered world of biology that allows different kinds of cells to synthesize protein at different rates and in different ways, and that those differences are important for cellular survival.”
In a previous study, researchers discovered that, by modifying the antibiotic puromycin, they could measure protein synthesis in rare cells in vivo.
Dr Morrison and his colleagues realized they could adapt this reagent to measure protein synthesis in HSCs and other cells in the hematopoietic system.
Their analyses showed that different types of blood cells produced vastly different amounts of protein per hour. And HSCs, in particular, synthesized much less protein than other hematopoietic progenitors.
“This result suggests that blood-forming stem cells require a lower rate of protein synthesis as compared to other blood-forming cells,” Dr Morrison said.
He and his colleagues then generated mice with a mutation in a component of the ribosome (Rpl24Bst/+ mice). HSCs in these mice had a 30% lower rate of protein production than controls.
The researchers observed the opposite effect when they deleted the tumor suppressor gene Pten in mouse HSCs. These mice saw a roughly 30% increase in protein production relative to controls.
However, as in the Rpl24Bst/+ mice, HSC function was noticeably impaired in these animals.
Together, these observations suggest that HSCs require a highly regulated rate of protein synthesis, such that increases or decreases in that rate impair HSC function.
“Amazingly, when the ribosomal mutant mice and the Pten mutant mice were bred together, stem cell function returned to normal, and we greatly delayed, and in some instances entirely blocked, the development of leukemia,” Dr Morrison said.
“All of this happened because protein production in stem cells was returned to normal. It was as if two wrongs made a right.”
Warmer temperatures push malaria to higher elevations

Credit: Asnakew Yeshiwondim
Researchers say they have the first hard evidence that malaria creeps to higher elevations during warmer years and retreats to lower altitudes when temperatures cool.
The evidence comes from an analysis of highland regions in Ethiopia and Colombia.
It suggests that future climate warming will prompt a rise in malaria incidence in densely populated regions of Africa and
South America, unless efforts to monitor and control malaria are increased.
“We saw an upward expansion of malaria cases to higher altitudes in warmer years, which is a clear signal of a response by highland malaria to changes in climate,” said study author Mercedes Pascual, PhD, of the University of Michigan in Ann Arbor.
“This is indisputable evidence of a climate effect. The main implication is that, with warmer temperatures, we expect to see a higher number of people exposed to the risk of malaria in tropical highland areas like these.”
Dr Pascual and her colleagues reported these findings in Science.
It was more than 20 years ago that malaria was first identified as a disease that might be especially sensitive to climate change, because both the Plasmodium parasites that cause it and the Anopheles mosquitoes that spread it thrive as temperatures warm.
Some early studies concluded that climate change would lead to an increase in malaria cases as the disease expanded its range into higher elevations. But some of the assumptions behind those predictions were later criticized.
More recently, researchers have argued that improved socioeconomic conditions and more aggressive mosquito-control efforts will likely exert a far greater influence than climatic factors over the extent and intensity of malaria worldwide.
What’s been missing in this debate is an analysis of regional records with sufficient resolution to determine how the spatial distribution of malaria cases has changed in response to year-to-year temperature variations, especially in densely populated highlands that have historically provided havens from the disease.
So Dr Pascual and her colleagues looked for evidence of a changing spatial distribution of malaria with varying temperature in the highlands of Ethiopia and Colombia. They examined malaria case records from the Antioquia region of western Colombia from 1990 to 2005 and from the Debre Zeit area of central Ethiopia from 1993 to 2005.
By focusing solely on the altitudinal response to year-to-year temperature changes, the researchers were able to exclude other variables that can influence malaria case numbers, such as mosquito-control programs, resistance to antimalarial drugs, and fluctuations in rainfall amounts.
The team found that the median altitude of malaria cases shifted to higher elevations in warmer years and back to lower elevations in cooler years. This relatively simple analysis yielded a clear signal that can only be explained by temperature changes, the group said.
“Our latest research suggests that, with progressive global warming, malaria will creep up the mountains and spread to new high-altitude areas,” said study author Menno Bouma, MD, of the London School of Hygiene & Tropical Medicine in the UK.
“And because these populations lack protective immunity, they will be particularly vulnerable to severe morbidity and mortality.”
In addition, the study results suggest that climate change can explain malaria trends in both the highland regions in recent decades.
In the Debre Zeit region of Ethiopia, at an elevation range of between 5280 feet and 7920 feet, about 37 million people (roughly 43% of the country’s population) live in rural areas at risk of higher malaria exposure under a warming climate.
In a previous study, researchers estimated that a 1-degree temperature increase could result in an additional 3 million malaria cases annually in Ethiopia in the under-15 population, unless control efforts are strengthened.
“Our findings here underscore the size of the problem,” Dr Pascual said, “and emphasize the need for sustained intervention efforts in these regions, especially in Africa.”

Credit: Asnakew Yeshiwondim
Researchers say they have the first hard evidence that malaria creeps to higher elevations during warmer years and retreats to lower altitudes when temperatures cool.
The evidence comes from an analysis of highland regions in Ethiopia and Colombia.
It suggests that future climate warming will prompt a rise in malaria incidence in densely populated regions of Africa and
South America, unless efforts to monitor and control malaria are increased.
“We saw an upward expansion of malaria cases to higher altitudes in warmer years, which is a clear signal of a response by highland malaria to changes in climate,” said study author Mercedes Pascual, PhD, of the University of Michigan in Ann Arbor.
“This is indisputable evidence of a climate effect. The main implication is that, with warmer temperatures, we expect to see a higher number of people exposed to the risk of malaria in tropical highland areas like these.”
Dr Pascual and her colleagues reported these findings in Science.
It was more than 20 years ago that malaria was first identified as a disease that might be especially sensitive to climate change, because both the Plasmodium parasites that cause it and the Anopheles mosquitoes that spread it thrive as temperatures warm.
Some early studies concluded that climate change would lead to an increase in malaria cases as the disease expanded its range into higher elevations. But some of the assumptions behind those predictions were later criticized.
More recently, researchers have argued that improved socioeconomic conditions and more aggressive mosquito-control efforts will likely exert a far greater influence than climatic factors over the extent and intensity of malaria worldwide.
What’s been missing in this debate is an analysis of regional records with sufficient resolution to determine how the spatial distribution of malaria cases has changed in response to year-to-year temperature variations, especially in densely populated highlands that have historically provided havens from the disease.
So Dr Pascual and her colleagues looked for evidence of a changing spatial distribution of malaria with varying temperature in the highlands of Ethiopia and Colombia. They examined malaria case records from the Antioquia region of western Colombia from 1990 to 2005 and from the Debre Zeit area of central Ethiopia from 1993 to 2005.
By focusing solely on the altitudinal response to year-to-year temperature changes, the researchers were able to exclude other variables that can influence malaria case numbers, such as mosquito-control programs, resistance to antimalarial drugs, and fluctuations in rainfall amounts.
The team found that the median altitude of malaria cases shifted to higher elevations in warmer years and back to lower elevations in cooler years. This relatively simple analysis yielded a clear signal that can only be explained by temperature changes, the group said.
“Our latest research suggests that, with progressive global warming, malaria will creep up the mountains and spread to new high-altitude areas,” said study author Menno Bouma, MD, of the London School of Hygiene & Tropical Medicine in the UK.
“And because these populations lack protective immunity, they will be particularly vulnerable to severe morbidity and mortality.”
In addition, the study results suggest that climate change can explain malaria trends in both the highland regions in recent decades.
In the Debre Zeit region of Ethiopia, at an elevation range of between 5280 feet and 7920 feet, about 37 million people (roughly 43% of the country’s population) live in rural areas at risk of higher malaria exposure under a warming climate.
In a previous study, researchers estimated that a 1-degree temperature increase could result in an additional 3 million malaria cases annually in Ethiopia in the under-15 population, unless control efforts are strengthened.
“Our findings here underscore the size of the problem,” Dr Pascual said, “and emphasize the need for sustained intervention efforts in these regions, especially in Africa.”

Credit: Asnakew Yeshiwondim
Researchers say they have the first hard evidence that malaria creeps to higher elevations during warmer years and retreats to lower altitudes when temperatures cool.
The evidence comes from an analysis of highland regions in Ethiopia and Colombia.
It suggests that future climate warming will prompt a rise in malaria incidence in densely populated regions of Africa and
South America, unless efforts to monitor and control malaria are increased.
“We saw an upward expansion of malaria cases to higher altitudes in warmer years, which is a clear signal of a response by highland malaria to changes in climate,” said study author Mercedes Pascual, PhD, of the University of Michigan in Ann Arbor.
“This is indisputable evidence of a climate effect. The main implication is that, with warmer temperatures, we expect to see a higher number of people exposed to the risk of malaria in tropical highland areas like these.”
Dr Pascual and her colleagues reported these findings in Science.
It was more than 20 years ago that malaria was first identified as a disease that might be especially sensitive to climate change, because both the Plasmodium parasites that cause it and the Anopheles mosquitoes that spread it thrive as temperatures warm.
Some early studies concluded that climate change would lead to an increase in malaria cases as the disease expanded its range into higher elevations. But some of the assumptions behind those predictions were later criticized.
More recently, researchers have argued that improved socioeconomic conditions and more aggressive mosquito-control efforts will likely exert a far greater influence than climatic factors over the extent and intensity of malaria worldwide.
What’s been missing in this debate is an analysis of regional records with sufficient resolution to determine how the spatial distribution of malaria cases has changed in response to year-to-year temperature variations, especially in densely populated highlands that have historically provided havens from the disease.
So Dr Pascual and her colleagues looked for evidence of a changing spatial distribution of malaria with varying temperature in the highlands of Ethiopia and Colombia. They examined malaria case records from the Antioquia region of western Colombia from 1990 to 2005 and from the Debre Zeit area of central Ethiopia from 1993 to 2005.
By focusing solely on the altitudinal response to year-to-year temperature changes, the researchers were able to exclude other variables that can influence malaria case numbers, such as mosquito-control programs, resistance to antimalarial drugs, and fluctuations in rainfall amounts.
The team found that the median altitude of malaria cases shifted to higher elevations in warmer years and back to lower elevations in cooler years. This relatively simple analysis yielded a clear signal that can only be explained by temperature changes, the group said.
“Our latest research suggests that, with progressive global warming, malaria will creep up the mountains and spread to new high-altitude areas,” said study author Menno Bouma, MD, of the London School of Hygiene & Tropical Medicine in the UK.
“And because these populations lack protective immunity, they will be particularly vulnerable to severe morbidity and mortality.”
In addition, the study results suggest that climate change can explain malaria trends in both the highland regions in recent decades.
In the Debre Zeit region of Ethiopia, at an elevation range of between 5280 feet and 7920 feet, about 37 million people (roughly 43% of the country’s population) live in rural areas at risk of higher malaria exposure under a warming climate.
In a previous study, researchers estimated that a 1-degree temperature increase could result in an additional 3 million malaria cases annually in Ethiopia in the under-15 population, unless control efforts are strengthened.
“Our findings here underscore the size of the problem,” Dr Pascual said, “and emphasize the need for sustained intervention efforts in these regions, especially in Africa.”
England’s Cancer Drugs Fund raises concerns

Credit: Rhoda Baer
Cancer patients in England are more likely to receive prescriptions for expensive drugs than patients in Wales, according to a study published in the British Journal of Cancer.
The research suggests this disparity is associated with the Cancer Drugs Fund (CDF), money set aside by the English government to pay for drugs that haven’t been approved by the National Institute for Health and Care Excellence (NICE) and aren’t available within the National Health Service (NHS).
The governments of Wales, Scotland, and Northern Ireland do not have access to the CDF or have similar programs of their own.
“There’s been much debate surrounding the Cancer Drugs Fund,” said study author Charlotte Chamberlain, MBBS, of the University of Bristol in the UK.
“The vast majority of Cancer Drugs Fund drugs do not cure the cancer but may extend life or improve symptoms in some people. The high cost of these drugs means that the NHS cannot afford other treatments, and, therefore, critics argue that public money is being spent inefficiently.”
To assess the impact of the CDF, Dr Chamberlain and her colleagues analyzed data from hospital pharmacies in England and Wales from August 2007 to December 2012. (The CDF was established in 2010, and the researchers wanted to capture data from before and after its introduction.)
The team evaluated 15 drugs that represent different categories of NICE approval—recommended, not recommended, and not yet appraised.
The results showed that, after the CDF was established, drugs recommended by NICE were not prescribed any more in England than in Wales.
However, drugs that were rejected by NICE because they were not cost-effective were prescribed up to 7 times more often in England than in Wales. For example, in the year before the CDF was introduced, prescription rates of imatinib (which is not recommended by NICE) were substantially higher in England than in Wales.
Immediately before the introduction of the CDF, following the first NICE rejection, imatinib prescribing declined in both countries. But it declined more slowly in England than in Wales, despite 2 additional NICE rejections. Regression analysis showed evidence of an association between the CDF and increased prescribing in England compared to Wales (P<0.001).
The research also revealed surprising information regarding the 3 most recently launched drugs—bendamustine, pazopanib, and abiraterone, which were awaiting NICE appraisal when the CDF was established but have since been approved.
These drugs were prescribed less often in England than in Wales. For instance, prescription rates of bendamustine were 25% lower in England.
This finding suggests that physicians in England have been slower to adopt newer drugs that are cost-effective, the researchers said.
“Our research has highlighted that the CDF has created an inequality between cancer sufferers in England and those in Wales,” Dr Chamberlain noted. “This raises ethical, moral, financial, and policy concerns.”

Credit: Rhoda Baer
Cancer patients in England are more likely to receive prescriptions for expensive drugs than patients in Wales, according to a study published in the British Journal of Cancer.
The research suggests this disparity is associated with the Cancer Drugs Fund (CDF), money set aside by the English government to pay for drugs that haven’t been approved by the National Institute for Health and Care Excellence (NICE) and aren’t available within the National Health Service (NHS).
The governments of Wales, Scotland, and Northern Ireland do not have access to the CDF or have similar programs of their own.
“There’s been much debate surrounding the Cancer Drugs Fund,” said study author Charlotte Chamberlain, MBBS, of the University of Bristol in the UK.
“The vast majority of Cancer Drugs Fund drugs do not cure the cancer but may extend life or improve symptoms in some people. The high cost of these drugs means that the NHS cannot afford other treatments, and, therefore, critics argue that public money is being spent inefficiently.”
To assess the impact of the CDF, Dr Chamberlain and her colleagues analyzed data from hospital pharmacies in England and Wales from August 2007 to December 2012. (The CDF was established in 2010, and the researchers wanted to capture data from before and after its introduction.)
The team evaluated 15 drugs that represent different categories of NICE approval—recommended, not recommended, and not yet appraised.
The results showed that, after the CDF was established, drugs recommended by NICE were not prescribed any more in England than in Wales.
However, drugs that were rejected by NICE because they were not cost-effective were prescribed up to 7 times more often in England than in Wales. For example, in the year before the CDF was introduced, prescription rates of imatinib (which is not recommended by NICE) were substantially higher in England than in Wales.
Immediately before the introduction of the CDF, following the first NICE rejection, imatinib prescribing declined in both countries. But it declined more slowly in England than in Wales, despite 2 additional NICE rejections. Regression analysis showed evidence of an association between the CDF and increased prescribing in England compared to Wales (P<0.001).
The research also revealed surprising information regarding the 3 most recently launched drugs—bendamustine, pazopanib, and abiraterone, which were awaiting NICE appraisal when the CDF was established but have since been approved.
These drugs were prescribed less often in England than in Wales. For instance, prescription rates of bendamustine were 25% lower in England.
This finding suggests that physicians in England have been slower to adopt newer drugs that are cost-effective, the researchers said.
“Our research has highlighted that the CDF has created an inequality between cancer sufferers in England and those in Wales,” Dr Chamberlain noted. “This raises ethical, moral, financial, and policy concerns.”

Credit: Rhoda Baer
Cancer patients in England are more likely to receive prescriptions for expensive drugs than patients in Wales, according to a study published in the British Journal of Cancer.
The research suggests this disparity is associated with the Cancer Drugs Fund (CDF), money set aside by the English government to pay for drugs that haven’t been approved by the National Institute for Health and Care Excellence (NICE) and aren’t available within the National Health Service (NHS).
The governments of Wales, Scotland, and Northern Ireland do not have access to the CDF or have similar programs of their own.
“There’s been much debate surrounding the Cancer Drugs Fund,” said study author Charlotte Chamberlain, MBBS, of the University of Bristol in the UK.
“The vast majority of Cancer Drugs Fund drugs do not cure the cancer but may extend life or improve symptoms in some people. The high cost of these drugs means that the NHS cannot afford other treatments, and, therefore, critics argue that public money is being spent inefficiently.”
To assess the impact of the CDF, Dr Chamberlain and her colleagues analyzed data from hospital pharmacies in England and Wales from August 2007 to December 2012. (The CDF was established in 2010, and the researchers wanted to capture data from before and after its introduction.)
The team evaluated 15 drugs that represent different categories of NICE approval—recommended, not recommended, and not yet appraised.
The results showed that, after the CDF was established, drugs recommended by NICE were not prescribed any more in England than in Wales.
However, drugs that were rejected by NICE because they were not cost-effective were prescribed up to 7 times more often in England than in Wales. For example, in the year before the CDF was introduced, prescription rates of imatinib (which is not recommended by NICE) were substantially higher in England than in Wales.
Immediately before the introduction of the CDF, following the first NICE rejection, imatinib prescribing declined in both countries. But it declined more slowly in England than in Wales, despite 2 additional NICE rejections. Regression analysis showed evidence of an association between the CDF and increased prescribing in England compared to Wales (P<0.001).
The research also revealed surprising information regarding the 3 most recently launched drugs—bendamustine, pazopanib, and abiraterone, which were awaiting NICE appraisal when the CDF was established but have since been approved.
These drugs were prescribed less often in England than in Wales. For instance, prescription rates of bendamustine were 25% lower in England.
This finding suggests that physicians in England have been slower to adopt newer drugs that are cost-effective, the researchers said.
“Our research has highlighted that the CDF has created an inequality between cancer sufferers in England and those in Wales,” Dr Chamberlain noted. “This raises ethical, moral, financial, and policy concerns.”