User login
Palliative chemo can have undesired outcomes
Credit: Rhoda Baer
Palliative chemotherapy can negatively impact the end of life for terminally ill cancer patients, according to a paper published in BMJ.
Investigators found that patients who received palliative chemotherapy in their last months of life had an increased risk of requiring intensive medical care, such as resuscitation, and dying in a place they did not choose, such as an intensive care unit.
The researchers therefore suggested that end-of-life discussions may be particularly important for patients who want to receive palliative chemotherapy.
“The results highlight the need for more effective communication by doctors of terminal prognoses and the likely outcomes of chemotherapy for these patients,” said study author Holly Prigerson, PhD, of Weill Cornell Medical College in New York.
“For patients to make informed choices about their care, they need to know if they are incurable and understand what their life expectancy is, that palliative chemotherapy is not intended to cure them, that it may not appreciably prolong their life, and that it may result in the receipt of very aggressive, life-prolonging care at the expense of their quality of life.”
Data have suggested that between 20% and 50% of patients with incurable cancers undergo palliative chemotherapy within 30 days of death. But it has not been clear whether the use of chemotherapy in a patient’s last months is associated with the need for intensive medical care in the last week of life or with the patient’s death.
So Dr Prigerson and her colleagues decided to study the use of palliative chemotherapy in patients with 6 or fewer months to live. The researchers used data from “Coping with Cancer,” a 6-year study of 386 terminally ill patients.
The patients were interviewed around the time of their decision regarding palliative chemotherapy. In the month after each patient died, caregivers were asked to rate their loved ones’ care, quality of life, and place of death as being where the patient would have wanted to die. The investigators then reviewed patients’ medical charts to determine the type of care they actually received in their last week.
Effects of palliative chemo
In all, 56% of patients opted to receive palliative chemotherapy. They were more likely to be younger, married, and better educated than patients not on the treatment.
Patients on chemotherapy also had better performance status, overall quality of life, physical functioning, and psychological well-being at study enrollment.
However, patients who received palliative chemotherapy had a greater risk of requiring cardiopulmonary resuscitation and/or mechanical ventilation (14% vs 2%), and they were more likely to need a feeding tube (11% vs 5%) in their last weeks of life.
Patients on chemotherapy had a greater risk of being admitted to an intensive care unit (14% vs 8%) and of having a late hospice referral (54% vs 37%).
They were also less likely to die where they wanted to (65% vs 80%). They had a greater risk of dying in an intensive care unit (11% vs 2%) and were less likely than their peers to die at home (47% vs 66%).
“It’s hard to see in these data much of a silver lining to palliative chemotherapy for patients in the terminal stage of their cancer,” Dr Prigerson said. “Until now, there hasn’t been evidence of harmful effects of palliative chemotherapy in the last few months of life.”
“This study is a first step in providing evidence that specifically demonstrates what negative outcomes may result. Additional studies are needed to confirm these troubling findings.”
Explaining the negative effects
Dr Prigerson said the harmful effects of palliative chemotherapy may be a result of misunderstanding, a lack of communication, and denial. Patients may not comprehend the purpose and likely consequences of palliative chemotherapy, and they may not fully acknowledge their own prognoses.
In the study, patients receiving palliative chemotherapy were less likely than their peers to talk to their oncologists about end-of-life care (37% vs 48%), to complete Do-Not-Resuscitate orders (36% vs 49%), or to acknowledge that they were terminally ill (35% vs 47%).
“Our finding that patients with terminal cancers were at higher risk of receiving intensive end-of-life care if they were treated with palliative chemotherapy months earlier underscores the importance of oncologists asking patients about their end-of-life wishes,” said Alexi Wright, MD, of the Dana-Farber Cancer Institute in Boston.
“We often wait until patients stop chemotherapy before asking them about where and how they want to die, but this study shows we need to ask patients about their preferences while they are receiving chemotherapy to ensure they receive the kind of care they want near death.”
Moving forward
The investigators stressed that the study results do not suggest patients should be denied palliative chemotherapy.
“The vast majority of patients in this study wanted palliative chemotherapy if it might increase their survival by as little as a week,” Dr Wright said. “This study is a step towards understanding some of the human costs and benefits of palliative chemotherapy.”
The researchers said additional studies should examine whether patients who are aware that chemotherapy is not intended to cure them still want to receive the treatment, confirm the negative outcomes of palliative chemotherapy, and determine if end-of-life discussions promote more informed decision-making and receipt of value-consistent care.
In a related editorial, Mike Rabow, MD, of the University of California, San Francisco, noted that although most patients with metastatic cancer choose to receive chemotherapy, evidence suggests most do not understand its intent.
He said Dr Prigerson’s study suggests the need to “better identify patients who are likely to benefit from chemotherapy near the end of life.” And he encouraged oncologists to discuss with patients the broader implications of chemotherapy when making decisions about treatment. ![]()

Credit: Rhoda Baer
Palliative chemotherapy can negatively impact the end of life for terminally ill cancer patients, according to a paper published in BMJ.
Investigators found that patients who received palliative chemotherapy in their last months of life had an increased risk of requiring intensive medical care, such as resuscitation, and dying in a place they did not choose, such as an intensive care unit.
The researchers therefore suggested that end-of-life discussions may be particularly important for patients who want to receive palliative chemotherapy.
“The results highlight the need for more effective communication by doctors of terminal prognoses and the likely outcomes of chemotherapy for these patients,” said study author Holly Prigerson, PhD, of Weill Cornell Medical College in New York.
“For patients to make informed choices about their care, they need to know if they are incurable and understand what their life expectancy is, that palliative chemotherapy is not intended to cure them, that it may not appreciably prolong their life, and that it may result in the receipt of very aggressive, life-prolonging care at the expense of their quality of life.”
Data have suggested that between 20% and 50% of patients with incurable cancers undergo palliative chemotherapy within 30 days of death. But it has not been clear whether the use of chemotherapy in a patient’s last months is associated with the need for intensive medical care in the last week of life or with the patient’s death.
So Dr Prigerson and her colleagues decided to study the use of palliative chemotherapy in patients with 6 or fewer months to live. The researchers used data from “Coping with Cancer,” a 6-year study of 386 terminally ill patients.
The patients were interviewed around the time of their decision regarding palliative chemotherapy. In the month after each patient died, caregivers were asked to rate their loved ones’ care, quality of life, and place of death as being where the patient would have wanted to die. The investigators then reviewed patients’ medical charts to determine the type of care they actually received in their last week.
Effects of palliative chemo
In all, 56% of patients opted to receive palliative chemotherapy. They were more likely to be younger, married, and better educated than patients not on the treatment.
Patients on chemotherapy also had better performance status, overall quality of life, physical functioning, and psychological well-being at study enrollment.
However, patients who received palliative chemotherapy had a greater risk of requiring cardiopulmonary resuscitation and/or mechanical ventilation (14% vs 2%), and they were more likely to need a feeding tube (11% vs 5%) in their last weeks of life.
Patients on chemotherapy had a greater risk of being admitted to an intensive care unit (14% vs 8%) and of having a late hospice referral (54% vs 37%).
They were also less likely to die where they wanted to (65% vs 80%). They had a greater risk of dying in an intensive care unit (11% vs 2%) and were less likely than their peers to die at home (47% vs 66%).
“It’s hard to see in these data much of a silver lining to palliative chemotherapy for patients in the terminal stage of their cancer,” Dr Prigerson said. “Until now, there hasn’t been evidence of harmful effects of palliative chemotherapy in the last few months of life.”
“This study is a first step in providing evidence that specifically demonstrates what negative outcomes may result. Additional studies are needed to confirm these troubling findings.”
Explaining the negative effects
Dr Prigerson said the harmful effects of palliative chemotherapy may be a result of misunderstanding, a lack of communication, and denial. Patients may not comprehend the purpose and likely consequences of palliative chemotherapy, and they may not fully acknowledge their own prognoses.
In the study, patients receiving palliative chemotherapy were less likely than their peers to talk to their oncologists about end-of-life care (37% vs 48%), to complete Do-Not-Resuscitate orders (36% vs 49%), or to acknowledge that they were terminally ill (35% vs 47%).
“Our finding that patients with terminal cancers were at higher risk of receiving intensive end-of-life care if they were treated with palliative chemotherapy months earlier underscores the importance of oncologists asking patients about their end-of-life wishes,” said Alexi Wright, MD, of the Dana-Farber Cancer Institute in Boston.
“We often wait until patients stop chemotherapy before asking them about where and how they want to die, but this study shows we need to ask patients about their preferences while they are receiving chemotherapy to ensure they receive the kind of care they want near death.”
Moving forward
The investigators stressed that the study results do not suggest patients should be denied palliative chemotherapy.
“The vast majority of patients in this study wanted palliative chemotherapy if it might increase their survival by as little as a week,” Dr Wright said. “This study is a step towards understanding some of the human costs and benefits of palliative chemotherapy.”
The researchers said additional studies should examine whether patients who are aware that chemotherapy is not intended to cure them still want to receive the treatment, confirm the negative outcomes of palliative chemotherapy, and determine if end-of-life discussions promote more informed decision-making and receipt of value-consistent care.
In a related editorial, Mike Rabow, MD, of the University of California, San Francisco, noted that although most patients with metastatic cancer choose to receive chemotherapy, evidence suggests most do not understand its intent.
He said Dr Prigerson’s study suggests the need to “better identify patients who are likely to benefit from chemotherapy near the end of life.” And he encouraged oncologists to discuss with patients the broader implications of chemotherapy when making decisions about treatment. ![]()

Credit: Rhoda Baer
Palliative chemotherapy can negatively impact the end of life for terminally ill cancer patients, according to a paper published in BMJ.
Investigators found that patients who received palliative chemotherapy in their last months of life had an increased risk of requiring intensive medical care, such as resuscitation, and dying in a place they did not choose, such as an intensive care unit.
The researchers therefore suggested that end-of-life discussions may be particularly important for patients who want to receive palliative chemotherapy.
“The results highlight the need for more effective communication by doctors of terminal prognoses and the likely outcomes of chemotherapy for these patients,” said study author Holly Prigerson, PhD, of Weill Cornell Medical College in New York.
“For patients to make informed choices about their care, they need to know if they are incurable and understand what their life expectancy is, that palliative chemotherapy is not intended to cure them, that it may not appreciably prolong their life, and that it may result in the receipt of very aggressive, life-prolonging care at the expense of their quality of life.”
Data have suggested that between 20% and 50% of patients with incurable cancers undergo palliative chemotherapy within 30 days of death. But it has not been clear whether the use of chemotherapy in a patient’s last months is associated with the need for intensive medical care in the last week of life or with the patient’s death.
So Dr Prigerson and her colleagues decided to study the use of palliative chemotherapy in patients with 6 or fewer months to live. The researchers used data from “Coping with Cancer,” a 6-year study of 386 terminally ill patients.
The patients were interviewed around the time of their decision regarding palliative chemotherapy. In the month after each patient died, caregivers were asked to rate their loved ones’ care, quality of life, and place of death as being where the patient would have wanted to die. The investigators then reviewed patients’ medical charts to determine the type of care they actually received in their last week.
Effects of palliative chemo
In all, 56% of patients opted to receive palliative chemotherapy. They were more likely to be younger, married, and better educated than patients not on the treatment.
Patients on chemotherapy also had better performance status, overall quality of life, physical functioning, and psychological well-being at study enrollment.
However, patients who received palliative chemotherapy had a greater risk of requiring cardiopulmonary resuscitation and/or mechanical ventilation (14% vs 2%), and they were more likely to need a feeding tube (11% vs 5%) in their last weeks of life.
Patients on chemotherapy had a greater risk of being admitted to an intensive care unit (14% vs 8%) and of having a late hospice referral (54% vs 37%).
They were also less likely to die where they wanted to (65% vs 80%). They had a greater risk of dying in an intensive care unit (11% vs 2%) and were less likely than their peers to die at home (47% vs 66%).
“It’s hard to see in these data much of a silver lining to palliative chemotherapy for patients in the terminal stage of their cancer,” Dr Prigerson said. “Until now, there hasn’t been evidence of harmful effects of palliative chemotherapy in the last few months of life.”
“This study is a first step in providing evidence that specifically demonstrates what negative outcomes may result. Additional studies are needed to confirm these troubling findings.”
Explaining the negative effects
Dr Prigerson said the harmful effects of palliative chemotherapy may be a result of misunderstanding, a lack of communication, and denial. Patients may not comprehend the purpose and likely consequences of palliative chemotherapy, and they may not fully acknowledge their own prognoses.
In the study, patients receiving palliative chemotherapy were less likely than their peers to talk to their oncologists about end-of-life care (37% vs 48%), to complete Do-Not-Resuscitate orders (36% vs 49%), or to acknowledge that they were terminally ill (35% vs 47%).
“Our finding that patients with terminal cancers were at higher risk of receiving intensive end-of-life care if they were treated with palliative chemotherapy months earlier underscores the importance of oncologists asking patients about their end-of-life wishes,” said Alexi Wright, MD, of the Dana-Farber Cancer Institute in Boston.
“We often wait until patients stop chemotherapy before asking them about where and how they want to die, but this study shows we need to ask patients about their preferences while they are receiving chemotherapy to ensure they receive the kind of care they want near death.”
Moving forward
The investigators stressed that the study results do not suggest patients should be denied palliative chemotherapy.
“The vast majority of patients in this study wanted palliative chemotherapy if it might increase their survival by as little as a week,” Dr Wright said. “This study is a step towards understanding some of the human costs and benefits of palliative chemotherapy.”
The researchers said additional studies should examine whether patients who are aware that chemotherapy is not intended to cure them still want to receive the treatment, confirm the negative outcomes of palliative chemotherapy, and determine if end-of-life discussions promote more informed decision-making and receipt of value-consistent care.
In a related editorial, Mike Rabow, MD, of the University of California, San Francisco, noted that although most patients with metastatic cancer choose to receive chemotherapy, evidence suggests most do not understand its intent.
He said Dr Prigerson’s study suggests the need to “better identify patients who are likely to benefit from chemotherapy near the end of life.” And he encouraged oncologists to discuss with patients the broader implications of chemotherapy when making decisions about treatment. ![]()
Histones’ role in gene regulation
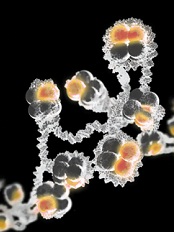
Credit: Eric Smith
Researchers say they’ve discovered how histones control PARP1’s ability to activate genes and repair DNA damage.
Their findings, published in Molecular Cell, appear to have implications for cancer treatment.
Specifically, the investigators found that chemical modification of the histone H2Av leads to substantial changes in nucleosome shape.
As a consequence, a previously hidden portion of the nucleosome becomes exposed and activates PARP1.
Upon activation, PARP1 assembles long branching molecules of Poly(ADP-ribose), which appear to open the DNA packaging around the site of PARP1 activation, thereby exposing specific genes for activation.
“[T]he nucleosome is often portrayed as a stable, inert structure, or a tiny ball,” said study author Alexei V. Tulin, PhD, of Fox Chase Cancer Center in Philadelphia.
“We found that the nucleosome is actually a quite dynamic structure. When we modified one histone, we changed the whole nucleosome.”
In addition to revealing new information about how histones control gene activation, Dr Tulin’s research elucidated a new mechanism of PARP1 regulation.
“This mechanism of PARP1 regulation by histones is still very new,” Dr Tulin said. “People believe that PARP1 is mainly activated through interactions with DNA, but we have found that the main pathway of PARP1 activation is through interactions with the nucleosome.”
Previous research suggested that combining standard anticancer agents with drugs that inhibit PARP1 can more effectively kill cancer cells. But clinical trials testing PARP1 inhibitors in cancer patients have produced disappointing results.
“I believe that, to a large extent, the previous setbacks were caused by a general misconception of the role of PARP1 in living cells and the mechanisms of PARP1 regulation,” Dr Tulin said. “Now that we know this mechanism of PARP1 regulation, we can design approaches to inhibit this protein in an effective way to better treat cancer.”
Dr Tulin and his colleagues are now developing the next generation of PARP1 inhibitors. Designed to block the newly identified mechanism of PARP1 activation, these inhibitors will specifically target PARP1, in contrast to the PARP1 inhibitors currently being tested in clinical trials. ![]()

Credit: Eric Smith
Researchers say they’ve discovered how histones control PARP1’s ability to activate genes and repair DNA damage.
Their findings, published in Molecular Cell, appear to have implications for cancer treatment.
Specifically, the investigators found that chemical modification of the histone H2Av leads to substantial changes in nucleosome shape.
As a consequence, a previously hidden portion of the nucleosome becomes exposed and activates PARP1.
Upon activation, PARP1 assembles long branching molecules of Poly(ADP-ribose), which appear to open the DNA packaging around the site of PARP1 activation, thereby exposing specific genes for activation.
“[T]he nucleosome is often portrayed as a stable, inert structure, or a tiny ball,” said study author Alexei V. Tulin, PhD, of Fox Chase Cancer Center in Philadelphia.
“We found that the nucleosome is actually a quite dynamic structure. When we modified one histone, we changed the whole nucleosome.”
In addition to revealing new information about how histones control gene activation, Dr Tulin’s research elucidated a new mechanism of PARP1 regulation.
“This mechanism of PARP1 regulation by histones is still very new,” Dr Tulin said. “People believe that PARP1 is mainly activated through interactions with DNA, but we have found that the main pathway of PARP1 activation is through interactions with the nucleosome.”
Previous research suggested that combining standard anticancer agents with drugs that inhibit PARP1 can more effectively kill cancer cells. But clinical trials testing PARP1 inhibitors in cancer patients have produced disappointing results.
“I believe that, to a large extent, the previous setbacks were caused by a general misconception of the role of PARP1 in living cells and the mechanisms of PARP1 regulation,” Dr Tulin said. “Now that we know this mechanism of PARP1 regulation, we can design approaches to inhibit this protein in an effective way to better treat cancer.”
Dr Tulin and his colleagues are now developing the next generation of PARP1 inhibitors. Designed to block the newly identified mechanism of PARP1 activation, these inhibitors will specifically target PARP1, in contrast to the PARP1 inhibitors currently being tested in clinical trials. ![]()

Credit: Eric Smith
Researchers say they’ve discovered how histones control PARP1’s ability to activate genes and repair DNA damage.
Their findings, published in Molecular Cell, appear to have implications for cancer treatment.
Specifically, the investigators found that chemical modification of the histone H2Av leads to substantial changes in nucleosome shape.
As a consequence, a previously hidden portion of the nucleosome becomes exposed and activates PARP1.
Upon activation, PARP1 assembles long branching molecules of Poly(ADP-ribose), which appear to open the DNA packaging around the site of PARP1 activation, thereby exposing specific genes for activation.
“[T]he nucleosome is often portrayed as a stable, inert structure, or a tiny ball,” said study author Alexei V. Tulin, PhD, of Fox Chase Cancer Center in Philadelphia.
“We found that the nucleosome is actually a quite dynamic structure. When we modified one histone, we changed the whole nucleosome.”
In addition to revealing new information about how histones control gene activation, Dr Tulin’s research elucidated a new mechanism of PARP1 regulation.
“This mechanism of PARP1 regulation by histones is still very new,” Dr Tulin said. “People believe that PARP1 is mainly activated through interactions with DNA, but we have found that the main pathway of PARP1 activation is through interactions with the nucleosome.”
Previous research suggested that combining standard anticancer agents with drugs that inhibit PARP1 can more effectively kill cancer cells. But clinical trials testing PARP1 inhibitors in cancer patients have produced disappointing results.
“I believe that, to a large extent, the previous setbacks were caused by a general misconception of the role of PARP1 in living cells and the mechanisms of PARP1 regulation,” Dr Tulin said. “Now that we know this mechanism of PARP1 regulation, we can design approaches to inhibit this protein in an effective way to better treat cancer.”
Dr Tulin and his colleagues are now developing the next generation of PARP1 inhibitors. Designed to block the newly identified mechanism of PARP1 activation, these inhibitors will specifically target PARP1, in contrast to the PARP1 inhibitors currently being tested in clinical trials. ![]()
Mutation responsible for insecticide resistance

spray insecticide
Credit: Morgana Wingard
A single genetic mutation can cause resistance to the main insecticides used to combat malaria, according to a study published in Genome Biology.
Researchers identified a mutation in the gene GSTe2 that allows mosquitoes to break down the insecticide DDT into non-toxic substances.
The mutation also makes mosquitoes resistant to pyrethroids, an insecticide class used in mosquito nets.
“We found a population of mosquitoes fully resistant to DDT but also to pyrethroids,” said study author Charles Wondji, PhD, of the Liverpool School of Tropical Medicine in the UK.
“So we wanted to elucidate the molecular basis of that resistance in the population and design a field-applicable diagnostic assay for its monitoring.”
To that end, the researchers did a genome-wide comparison on mosquitoes that were fully susceptible to insecticides and Anopheles funestus mosquitoes from the Republic of Benin in Africa, which were resistant to DDT and the pyrethroid permethrin.
The team found the GSTe2 gene was upregulated in the resistant mosquitoes. And a single mutation (L119F) changed a non-resistant version of the gene to an insecticide-resistant version.
The researchers then designed a DNA-based diagnostic test for this metabolic resistance and confirmed that this mutation was found in mosquitoes from other areas of the world with DDT resistance, but it was completely absent in regions without resistance.
X-ray crystallography of the protein coded by the gene illustrated exactly how the mutation conferred resistance—by opening up the active site where DDT molecules bind to the protein so that more can be broken down. In other words, the mosquito can survive by breaking down the poison into non-toxic substances.
The researchers also introduced the gene into Drosophila melanogaster and found the flies became resistant to DDT and pyrethroids, whereas control flies did not. The team said this confirms that a single mutation is enough to make insects resistant to both DDT and pyrethroids.
“For the first time, we have been able to identify a molecular marker for metabolic resistance in a mosquito population and to design a DNA-based diagnostic assay,” Dr Wondji said.
“Such tools will allow control programs to detect and track resistance at an early stage in the field, which is an essential requirement to successfully tackle the growing problem of insecticide resistance in vector control. This significant progress opens the door for us to do this with other forms of resistance as well and in other vector species.” ![]()

spray insecticide
Credit: Morgana Wingard
A single genetic mutation can cause resistance to the main insecticides used to combat malaria, according to a study published in Genome Biology.
Researchers identified a mutation in the gene GSTe2 that allows mosquitoes to break down the insecticide DDT into non-toxic substances.
The mutation also makes mosquitoes resistant to pyrethroids, an insecticide class used in mosquito nets.
“We found a population of mosquitoes fully resistant to DDT but also to pyrethroids,” said study author Charles Wondji, PhD, of the Liverpool School of Tropical Medicine in the UK.
“So we wanted to elucidate the molecular basis of that resistance in the population and design a field-applicable diagnostic assay for its monitoring.”
To that end, the researchers did a genome-wide comparison on mosquitoes that were fully susceptible to insecticides and Anopheles funestus mosquitoes from the Republic of Benin in Africa, which were resistant to DDT and the pyrethroid permethrin.
The team found the GSTe2 gene was upregulated in the resistant mosquitoes. And a single mutation (L119F) changed a non-resistant version of the gene to an insecticide-resistant version.
The researchers then designed a DNA-based diagnostic test for this metabolic resistance and confirmed that this mutation was found in mosquitoes from other areas of the world with DDT resistance, but it was completely absent in regions without resistance.
X-ray crystallography of the protein coded by the gene illustrated exactly how the mutation conferred resistance—by opening up the active site where DDT molecules bind to the protein so that more can be broken down. In other words, the mosquito can survive by breaking down the poison into non-toxic substances.
The researchers also introduced the gene into Drosophila melanogaster and found the flies became resistant to DDT and pyrethroids, whereas control flies did not. The team said this confirms that a single mutation is enough to make insects resistant to both DDT and pyrethroids.
“For the first time, we have been able to identify a molecular marker for metabolic resistance in a mosquito population and to design a DNA-based diagnostic assay,” Dr Wondji said.
“Such tools will allow control programs to detect and track resistance at an early stage in the field, which is an essential requirement to successfully tackle the growing problem of insecticide resistance in vector control. This significant progress opens the door for us to do this with other forms of resistance as well and in other vector species.” ![]()

spray insecticide
Credit: Morgana Wingard
A single genetic mutation can cause resistance to the main insecticides used to combat malaria, according to a study published in Genome Biology.
Researchers identified a mutation in the gene GSTe2 that allows mosquitoes to break down the insecticide DDT into non-toxic substances.
The mutation also makes mosquitoes resistant to pyrethroids, an insecticide class used in mosquito nets.
“We found a population of mosquitoes fully resistant to DDT but also to pyrethroids,” said study author Charles Wondji, PhD, of the Liverpool School of Tropical Medicine in the UK.
“So we wanted to elucidate the molecular basis of that resistance in the population and design a field-applicable diagnostic assay for its monitoring.”
To that end, the researchers did a genome-wide comparison on mosquitoes that were fully susceptible to insecticides and Anopheles funestus mosquitoes from the Republic of Benin in Africa, which were resistant to DDT and the pyrethroid permethrin.
The team found the GSTe2 gene was upregulated in the resistant mosquitoes. And a single mutation (L119F) changed a non-resistant version of the gene to an insecticide-resistant version.
The researchers then designed a DNA-based diagnostic test for this metabolic resistance and confirmed that this mutation was found in mosquitoes from other areas of the world with DDT resistance, but it was completely absent in regions without resistance.
X-ray crystallography of the protein coded by the gene illustrated exactly how the mutation conferred resistance—by opening up the active site where DDT molecules bind to the protein so that more can be broken down. In other words, the mosquito can survive by breaking down the poison into non-toxic substances.
The researchers also introduced the gene into Drosophila melanogaster and found the flies became resistant to DDT and pyrethroids, whereas control flies did not. The team said this confirms that a single mutation is enough to make insects resistant to both DDT and pyrethroids.
“For the first time, we have been able to identify a molecular marker for metabolic resistance in a mosquito population and to design a DNA-based diagnostic assay,” Dr Wondji said.
“Such tools will allow control programs to detect and track resistance at an early stage in the field, which is an essential requirement to successfully tackle the growing problem of insecticide resistance in vector control. This significant progress opens the door for us to do this with other forms of resistance as well and in other vector species.” ![]()
Computerized checklist can reduce CLABSI rate
Staphylococcus infection
Credit: Bill Branson
A computerized safety checklist that pulls information from patients’ electronic medical records can reduce the incidence of central line-associated bloodstream infections (CLABSIs), according to a study published in Pediatrics.
The study was conducted among children admitted to the pediatric intensive care unit at Lucile Packard Children’s Hospital Stanford in California.
Researchers found the safety checklist increased overall staff compliance with best practices for CLABSI prevention and resulted in a 3-fold reduction in CLABSI incidence.
The automated checklist, and a dashboard-style interface used to interact with it, was designed to help caregivers follow national guidelines for CLABSI prevention. The system combed through data in a patient’s electronic medical record and pushed alerts to physicians and nurses when a patient’s central line was due for care.
The dashboard interface displayed real-time alerts on a large LCD screen in the nurses’ station. Alerts—shown as red, yellow, or green dots beside patients’ names—were generated if, for example, the dressing on a patient’s central line was due to be changed, or if it was time for caregivers to re-evaluate whether medications given in the central line could be switched to oral formulations instead.
“The information was visible and easy to digest,” said study author Deborah Franzon, MD. “We improved compliance with best-care practices and pulled information that otherwise would have been difficult to look for. It reduced busy work and made it possible for the healthcare team to perform their jobs more efficiently and effectively.”
The system was implemented on May 1, 2011, but the researchers considered the rollout period to extend to August 31, 2011. So this period was not included in the analysis.
The team compared data on CLABSI rates, compliance with bundle elements, and staff perceptions/knowledge before the intervention began—from June 1, 2009, to April 30, 2011—and after the system was fully implemented—September 1, 2011, to December 31, 2012.
CLABSI rates decreased from 2.6 per 1000 line-days before the intervention to 0.7 per 1000 line-days afterward (P=0.02). There were a total of 19 CLABSIs per 7322 line-days pre-intervention and 7 CLABSIs per 6155 line-days post-intervention.
The researchers estimated that the intervention saved approximately $260,000 per year in healthcare costs. Treating a single CLABSI costs approximately $39,000.
The team also found that daily documentation of line necessity increased from 30% before the intervention to 73% after (P<0.001). Compliance with dressing changes increased from 87% to 90% (P=0.003).
Compliance with cap changes increased from 87% to 93% (P<0.001). And compliance with port needle changes increased from 69% to 95% (P<0.001). However, compliance with insertion bundle documentation decreased from 67% to 62% (P=0.001).
After the system was implemented, there was a significant increase in staff perception that the medical team addressed central line necessity during rounds (P=0.02). But there was no significant difference in communication among team members (P=0.73) or knowledge regarding the components of the maintenance bundle (P=0.39).
Nevertheless, the researchers concluded that their system promotes compliance with best practices for CLABSI prevention, thereby reducing the risk of harm to patients.
The team hopes to use the system in other ways, such as monitoring the recovery of children who have received organ transplants.
“[The system] lets physicians focus on taking care of the patient while automating some of the background safety checks,” said study author Natalie Pageler, MD. “The nice thing about this tool is that it’s integrated into the electronic medical record, which we use every single day.” ![]()

Staphylococcus infection
Credit: Bill Branson
A computerized safety checklist that pulls information from patients’ electronic medical records can reduce the incidence of central line-associated bloodstream infections (CLABSIs), according to a study published in Pediatrics.
The study was conducted among children admitted to the pediatric intensive care unit at Lucile Packard Children’s Hospital Stanford in California.
Researchers found the safety checklist increased overall staff compliance with best practices for CLABSI prevention and resulted in a 3-fold reduction in CLABSI incidence.
The automated checklist, and a dashboard-style interface used to interact with it, was designed to help caregivers follow national guidelines for CLABSI prevention. The system combed through data in a patient’s electronic medical record and pushed alerts to physicians and nurses when a patient’s central line was due for care.
The dashboard interface displayed real-time alerts on a large LCD screen in the nurses’ station. Alerts—shown as red, yellow, or green dots beside patients’ names—were generated if, for example, the dressing on a patient’s central line was due to be changed, or if it was time for caregivers to re-evaluate whether medications given in the central line could be switched to oral formulations instead.
“The information was visible and easy to digest,” said study author Deborah Franzon, MD. “We improved compliance with best-care practices and pulled information that otherwise would have been difficult to look for. It reduced busy work and made it possible for the healthcare team to perform their jobs more efficiently and effectively.”
The system was implemented on May 1, 2011, but the researchers considered the rollout period to extend to August 31, 2011. So this period was not included in the analysis.
The team compared data on CLABSI rates, compliance with bundle elements, and staff perceptions/knowledge before the intervention began—from June 1, 2009, to April 30, 2011—and after the system was fully implemented—September 1, 2011, to December 31, 2012.
CLABSI rates decreased from 2.6 per 1000 line-days before the intervention to 0.7 per 1000 line-days afterward (P=0.02). There were a total of 19 CLABSIs per 7322 line-days pre-intervention and 7 CLABSIs per 6155 line-days post-intervention.
The researchers estimated that the intervention saved approximately $260,000 per year in healthcare costs. Treating a single CLABSI costs approximately $39,000.
The team also found that daily documentation of line necessity increased from 30% before the intervention to 73% after (P<0.001). Compliance with dressing changes increased from 87% to 90% (P=0.003).
Compliance with cap changes increased from 87% to 93% (P<0.001). And compliance with port needle changes increased from 69% to 95% (P<0.001). However, compliance with insertion bundle documentation decreased from 67% to 62% (P=0.001).
After the system was implemented, there was a significant increase in staff perception that the medical team addressed central line necessity during rounds (P=0.02). But there was no significant difference in communication among team members (P=0.73) or knowledge regarding the components of the maintenance bundle (P=0.39).
Nevertheless, the researchers concluded that their system promotes compliance with best practices for CLABSI prevention, thereby reducing the risk of harm to patients.
The team hopes to use the system in other ways, such as monitoring the recovery of children who have received organ transplants.
“[The system] lets physicians focus on taking care of the patient while automating some of the background safety checks,” said study author Natalie Pageler, MD. “The nice thing about this tool is that it’s integrated into the electronic medical record, which we use every single day.” ![]()

Staphylococcus infection
Credit: Bill Branson
A computerized safety checklist that pulls information from patients’ electronic medical records can reduce the incidence of central line-associated bloodstream infections (CLABSIs), according to a study published in Pediatrics.
The study was conducted among children admitted to the pediatric intensive care unit at Lucile Packard Children’s Hospital Stanford in California.
Researchers found the safety checklist increased overall staff compliance with best practices for CLABSI prevention and resulted in a 3-fold reduction in CLABSI incidence.
The automated checklist, and a dashboard-style interface used to interact with it, was designed to help caregivers follow national guidelines for CLABSI prevention. The system combed through data in a patient’s electronic medical record and pushed alerts to physicians and nurses when a patient’s central line was due for care.
The dashboard interface displayed real-time alerts on a large LCD screen in the nurses’ station. Alerts—shown as red, yellow, or green dots beside patients’ names—were generated if, for example, the dressing on a patient’s central line was due to be changed, or if it was time for caregivers to re-evaluate whether medications given in the central line could be switched to oral formulations instead.
“The information was visible and easy to digest,” said study author Deborah Franzon, MD. “We improved compliance with best-care practices and pulled information that otherwise would have been difficult to look for. It reduced busy work and made it possible for the healthcare team to perform their jobs more efficiently and effectively.”
The system was implemented on May 1, 2011, but the researchers considered the rollout period to extend to August 31, 2011. So this period was not included in the analysis.
The team compared data on CLABSI rates, compliance with bundle elements, and staff perceptions/knowledge before the intervention began—from June 1, 2009, to April 30, 2011—and after the system was fully implemented—September 1, 2011, to December 31, 2012.
CLABSI rates decreased from 2.6 per 1000 line-days before the intervention to 0.7 per 1000 line-days afterward (P=0.02). There were a total of 19 CLABSIs per 7322 line-days pre-intervention and 7 CLABSIs per 6155 line-days post-intervention.
The researchers estimated that the intervention saved approximately $260,000 per year in healthcare costs. Treating a single CLABSI costs approximately $39,000.
The team also found that daily documentation of line necessity increased from 30% before the intervention to 73% after (P<0.001). Compliance with dressing changes increased from 87% to 90% (P=0.003).
Compliance with cap changes increased from 87% to 93% (P<0.001). And compliance with port needle changes increased from 69% to 95% (P<0.001). However, compliance with insertion bundle documentation decreased from 67% to 62% (P=0.001).
After the system was implemented, there was a significant increase in staff perception that the medical team addressed central line necessity during rounds (P=0.02). But there was no significant difference in communication among team members (P=0.73) or knowledge regarding the components of the maintenance bundle (P=0.39).
Nevertheless, the researchers concluded that their system promotes compliance with best practices for CLABSI prevention, thereby reducing the risk of harm to patients.
The team hopes to use the system in other ways, such as monitoring the recovery of children who have received organ transplants.
“[The system] lets physicians focus on taking care of the patient while automating some of the background safety checks,” said study author Natalie Pageler, MD. “The nice thing about this tool is that it’s integrated into the electronic medical record, which we use every single day.” ![]()
Malaria parasite originated in Africa, team says

David Morgan & Crickette Sanz
Investigators have found evidence suggesting the malaria parasite Plasmodium vivax originated in Africa.
Until recently, the closest genetic relatives of human P vivax were found only in Asian macaques, leading researchers to believe that P vivax originated in Asia.
The current study, published in Nature Communications, showed that wild apes in central Africa are widely infected with parasites that are, genetically, nearly identical to human P vivax.
This finding overturns the dogma that P vivax originated in Asia, despite being most prevalent in humans there now, and also solves other questions about P vivax infection.
For example, it explains why the Duffy-null phenotype, which confers resistance to P vivax, is common among people indigenous to Africa. And it explains how travelers returning from regions where most people are Duffy-negative can be infected with P vivax.
Paul Sharp, PhD, of the University of Edinburgh in the UK, and his colleagues conducted this research, testing more than 5000 ape fecal samples from dozens of field stations and sanctuaries in Africa for P vivax DNA.
They found P vivax-like sequences in chimpanzees, western gorillas, and eastern gorillas, but not in bonobos. Ape P vivax was highly prevalent in wild communities, exhibiting infection rates consistent with stable transmission of the parasite within the wild apes.
To examine the evolutionary relationships between ape and human parasites, the researchers generated parasite DNA sequences from wild and sanctuary apes, as well as from a global sampling of human P vivax infections.
They constructed a family tree of the sequences and found that ape and human parasites were very closely related. But ape parasites were more diverse than the human parasites and did not group according to their host species. The human parasites formed a single lineage that fell within the branches of ape parasite sequences.
From these evolutionary relationships, the investigators concluded that P vivax is of African—not Asian—origin and that all existing human P vivax parasites evolved from a single ancestor that spread out of Africa.
The high prevalence of P vivax in wild apes, along with the recent finding of ape P vivax in a European traveler, indicates the existence of a substantial natural reservoir of P vivax in Africa.
Resolving the Duffy-negative paradox
Of the 5 Plasmodium species known to cause malaria in humans, P vivax is the most widespread. Although highly prevalent in Asia and Latin America, P vivax was thought to be absent from west and central Africa due to a mutation that causes the Duffy-negative phenotype in most indigenous African people.
P vivax parasites enter human red blood cells via the Duffy protein receptor. Because the absence of the receptor on the surface of these cells confers protection against P vivax malaria, this parasite has long been suspected to be the agent that selected for this mutation. However, this hypothesis had been difficult to reconcile with the belief that P vivax originated in Asia.
“Our finding that wild-living apes in central Africa show widespread infection with diverse strains of P vivax provides new insight into the evolutionary history of human P vivax and resolves the paradox that a mutation conferring resistance to P vivax occurs with high frequency in the very region where this parasite is absent in humans,” said study author Beatrice Hahn, MD, of the University of Pennsylvania in Philadelphia.
“One interpretation of the relationships that we observed is that a single host switch from apes gave rise to human P vivax, analogous to the origin of human P falciparum,” Dr Sharp added. “However, this seems unlikely in this case, since ape P vivax does not divide into gorilla- and chimpanzee-specific lineages.”
A more plausible scenario, according to the researchers, is that an ancestral P vivax stock was able to infect humans, gorillas, and chimpanzees in Africa until the Duffy-negative mutation started to spread—around 30,000 years ago—and eliminated P vivax from humans there.
Under this scenario, existing human-infecting P vivax is a parasite that survived after spreading out of Africa.
“The existence of a P vivax reservoir within the forests of central Africa has public health implications,” said study author Martine Peeters, PhD, of the Institut de Recherche pour le Développement and the University of Montpellier in France.
“First, it solves the mystery of P vivax infections in travelers returning from regions where 99% of the human population is Duffy-negative. It also raises the possibility that Duffy-positive humans whose work may bring them in close proximity to chimpanzees and gorillas may become infected by ape P vivax. This has already happened once and may happen again, with unknown consequences.”
The investigators are also concerned about the possibility that ape P vivax may spread via international travel to countries where human P vivax is actively transmitted. Since ape P vivax is more genetically diverse than human P vivax, it may have more versatility to escape treatment and prevention measures, especially if human and ape parasites were able to recombine.
Given what biologists know about P vivax’s ability to switch hosts, the researchers suggest it is important to screen Duffy-positive and Duffy-negative humans in west central Africa, as well as transmitting mosquito vectors, for the presence of ape P vivax. The team believes this information is necessary to inform malaria control and eradication efforts of the propensity of ape P vivax to cross over to humans.
The investigators are also planning to compare and contrast the molecular and biological properties of human and ape parasites to identify host-specific interactions and transmission requirements, thereby uncovering vulnerabilities that can be exploited to combat human malaria. ![]()

David Morgan & Crickette Sanz
Investigators have found evidence suggesting the malaria parasite Plasmodium vivax originated in Africa.
Until recently, the closest genetic relatives of human P vivax were found only in Asian macaques, leading researchers to believe that P vivax originated in Asia.
The current study, published in Nature Communications, showed that wild apes in central Africa are widely infected with parasites that are, genetically, nearly identical to human P vivax.
This finding overturns the dogma that P vivax originated in Asia, despite being most prevalent in humans there now, and also solves other questions about P vivax infection.
For example, it explains why the Duffy-null phenotype, which confers resistance to P vivax, is common among people indigenous to Africa. And it explains how travelers returning from regions where most people are Duffy-negative can be infected with P vivax.
Paul Sharp, PhD, of the University of Edinburgh in the UK, and his colleagues conducted this research, testing more than 5000 ape fecal samples from dozens of field stations and sanctuaries in Africa for P vivax DNA.
They found P vivax-like sequences in chimpanzees, western gorillas, and eastern gorillas, but not in bonobos. Ape P vivax was highly prevalent in wild communities, exhibiting infection rates consistent with stable transmission of the parasite within the wild apes.
To examine the evolutionary relationships between ape and human parasites, the researchers generated parasite DNA sequences from wild and sanctuary apes, as well as from a global sampling of human P vivax infections.
They constructed a family tree of the sequences and found that ape and human parasites were very closely related. But ape parasites were more diverse than the human parasites and did not group according to their host species. The human parasites formed a single lineage that fell within the branches of ape parasite sequences.
From these evolutionary relationships, the investigators concluded that P vivax is of African—not Asian—origin and that all existing human P vivax parasites evolved from a single ancestor that spread out of Africa.
The high prevalence of P vivax in wild apes, along with the recent finding of ape P vivax in a European traveler, indicates the existence of a substantial natural reservoir of P vivax in Africa.
Resolving the Duffy-negative paradox
Of the 5 Plasmodium species known to cause malaria in humans, P vivax is the most widespread. Although highly prevalent in Asia and Latin America, P vivax was thought to be absent from west and central Africa due to a mutation that causes the Duffy-negative phenotype in most indigenous African people.
P vivax parasites enter human red blood cells via the Duffy protein receptor. Because the absence of the receptor on the surface of these cells confers protection against P vivax malaria, this parasite has long been suspected to be the agent that selected for this mutation. However, this hypothesis had been difficult to reconcile with the belief that P vivax originated in Asia.
“Our finding that wild-living apes in central Africa show widespread infection with diverse strains of P vivax provides new insight into the evolutionary history of human P vivax and resolves the paradox that a mutation conferring resistance to P vivax occurs with high frequency in the very region where this parasite is absent in humans,” said study author Beatrice Hahn, MD, of the University of Pennsylvania in Philadelphia.
“One interpretation of the relationships that we observed is that a single host switch from apes gave rise to human P vivax, analogous to the origin of human P falciparum,” Dr Sharp added. “However, this seems unlikely in this case, since ape P vivax does not divide into gorilla- and chimpanzee-specific lineages.”
A more plausible scenario, according to the researchers, is that an ancestral P vivax stock was able to infect humans, gorillas, and chimpanzees in Africa until the Duffy-negative mutation started to spread—around 30,000 years ago—and eliminated P vivax from humans there.
Under this scenario, existing human-infecting P vivax is a parasite that survived after spreading out of Africa.
“The existence of a P vivax reservoir within the forests of central Africa has public health implications,” said study author Martine Peeters, PhD, of the Institut de Recherche pour le Développement and the University of Montpellier in France.
“First, it solves the mystery of P vivax infections in travelers returning from regions where 99% of the human population is Duffy-negative. It also raises the possibility that Duffy-positive humans whose work may bring them in close proximity to chimpanzees and gorillas may become infected by ape P vivax. This has already happened once and may happen again, with unknown consequences.”
The investigators are also concerned about the possibility that ape P vivax may spread via international travel to countries where human P vivax is actively transmitted. Since ape P vivax is more genetically diverse than human P vivax, it may have more versatility to escape treatment and prevention measures, especially if human and ape parasites were able to recombine.
Given what biologists know about P vivax’s ability to switch hosts, the researchers suggest it is important to screen Duffy-positive and Duffy-negative humans in west central Africa, as well as transmitting mosquito vectors, for the presence of ape P vivax. The team believes this information is necessary to inform malaria control and eradication efforts of the propensity of ape P vivax to cross over to humans.
The investigators are also planning to compare and contrast the molecular and biological properties of human and ape parasites to identify host-specific interactions and transmission requirements, thereby uncovering vulnerabilities that can be exploited to combat human malaria. ![]()

David Morgan & Crickette Sanz
Investigators have found evidence suggesting the malaria parasite Plasmodium vivax originated in Africa.
Until recently, the closest genetic relatives of human P vivax were found only in Asian macaques, leading researchers to believe that P vivax originated in Asia.
The current study, published in Nature Communications, showed that wild apes in central Africa are widely infected with parasites that are, genetically, nearly identical to human P vivax.
This finding overturns the dogma that P vivax originated in Asia, despite being most prevalent in humans there now, and also solves other questions about P vivax infection.
For example, it explains why the Duffy-null phenotype, which confers resistance to P vivax, is common among people indigenous to Africa. And it explains how travelers returning from regions where most people are Duffy-negative can be infected with P vivax.
Paul Sharp, PhD, of the University of Edinburgh in the UK, and his colleagues conducted this research, testing more than 5000 ape fecal samples from dozens of field stations and sanctuaries in Africa for P vivax DNA.
They found P vivax-like sequences in chimpanzees, western gorillas, and eastern gorillas, but not in bonobos. Ape P vivax was highly prevalent in wild communities, exhibiting infection rates consistent with stable transmission of the parasite within the wild apes.
To examine the evolutionary relationships between ape and human parasites, the researchers generated parasite DNA sequences from wild and sanctuary apes, as well as from a global sampling of human P vivax infections.
They constructed a family tree of the sequences and found that ape and human parasites were very closely related. But ape parasites were more diverse than the human parasites and did not group according to their host species. The human parasites formed a single lineage that fell within the branches of ape parasite sequences.
From these evolutionary relationships, the investigators concluded that P vivax is of African—not Asian—origin and that all existing human P vivax parasites evolved from a single ancestor that spread out of Africa.
The high prevalence of P vivax in wild apes, along with the recent finding of ape P vivax in a European traveler, indicates the existence of a substantial natural reservoir of P vivax in Africa.
Resolving the Duffy-negative paradox
Of the 5 Plasmodium species known to cause malaria in humans, P vivax is the most widespread. Although highly prevalent in Asia and Latin America, P vivax was thought to be absent from west and central Africa due to a mutation that causes the Duffy-negative phenotype in most indigenous African people.
P vivax parasites enter human red blood cells via the Duffy protein receptor. Because the absence of the receptor on the surface of these cells confers protection against P vivax malaria, this parasite has long been suspected to be the agent that selected for this mutation. However, this hypothesis had been difficult to reconcile with the belief that P vivax originated in Asia.
“Our finding that wild-living apes in central Africa show widespread infection with diverse strains of P vivax provides new insight into the evolutionary history of human P vivax and resolves the paradox that a mutation conferring resistance to P vivax occurs with high frequency in the very region where this parasite is absent in humans,” said study author Beatrice Hahn, MD, of the University of Pennsylvania in Philadelphia.
“One interpretation of the relationships that we observed is that a single host switch from apes gave rise to human P vivax, analogous to the origin of human P falciparum,” Dr Sharp added. “However, this seems unlikely in this case, since ape P vivax does not divide into gorilla- and chimpanzee-specific lineages.”
A more plausible scenario, according to the researchers, is that an ancestral P vivax stock was able to infect humans, gorillas, and chimpanzees in Africa until the Duffy-negative mutation started to spread—around 30,000 years ago—and eliminated P vivax from humans there.
Under this scenario, existing human-infecting P vivax is a parasite that survived after spreading out of Africa.
“The existence of a P vivax reservoir within the forests of central Africa has public health implications,” said study author Martine Peeters, PhD, of the Institut de Recherche pour le Développement and the University of Montpellier in France.
“First, it solves the mystery of P vivax infections in travelers returning from regions where 99% of the human population is Duffy-negative. It also raises the possibility that Duffy-positive humans whose work may bring them in close proximity to chimpanzees and gorillas may become infected by ape P vivax. This has already happened once and may happen again, with unknown consequences.”
The investigators are also concerned about the possibility that ape P vivax may spread via international travel to countries where human P vivax is actively transmitted. Since ape P vivax is more genetically diverse than human P vivax, it may have more versatility to escape treatment and prevention measures, especially if human and ape parasites were able to recombine.
Given what biologists know about P vivax’s ability to switch hosts, the researchers suggest it is important to screen Duffy-positive and Duffy-negative humans in west central Africa, as well as transmitting mosquito vectors, for the presence of ape P vivax. The team believes this information is necessary to inform malaria control and eradication efforts of the propensity of ape P vivax to cross over to humans.
The investigators are also planning to compare and contrast the molecular and biological properties of human and ape parasites to identify host-specific interactions and transmission requirements, thereby uncovering vulnerabilities that can be exploited to combat human malaria. ![]()
Protein appears essential to malaria transmission
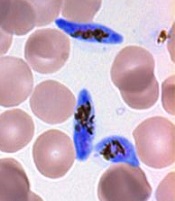
gametocyte stage (blue) and
uninfected red blood cells
Credit: The Llinás lab
Results of 2 new studies suggest that a single regulatory protein acts as a master switch to trigger development of the sexual forms of malaria parasites.
It appears that the protein, AP2-G, is necessary for activating a set of genes that initiate the development of Plasmodium gametocytes, the only forms of the parasite that are infectious to mosquitoes.
This suggests that if researchers can target AP2-G, they can stop sexual parasites from forming.
And if the sexual forms of the parasite never develop in an infected person’s blood, none will enter the mosquito’s gut, and the mosquito will be unable to infect anyone else with malaria.
“Exciting opportunities now lie ahead for finding an effective way to break the chain of malaria transmission by preventing the malaria parasite from completing its full lifecycle,” said Manuel Llinás, PhD, a professor at Pennsylvania State University who was involved in both studies.
The 2 studies, which were published as letters to Nature, had remarkably similar results, despite the fact that the groups worked with 2 different malaria parasites—Plasmodium falciparum and Plasmodium berghei.
In one study, researchers analyzed the whole-genome sequences of 2 P falciparum strains that were unable to produce gametocytes. The only mutated, non-functional gene common to both strains was the AP2-G gene.
In the other study, researchers sequenced P berghei parasites that had lost their ability to make gametocytes. Again, the only common mutated gene in these parasites was AP2-G.
To confirm these observations, both groups of researchers disabled the AP2-G gene in parasites that could generate gametocytes.
As expected, disabling the gene prevented the parasites from producing gametocytes. But the parasites regained their ability to make gametocytes when the mutated gene was repaired.
These results, as well as results of additional experiments, suggest that sexual-stage malaria parasites are produced only when the AP2-G protein is in working order.
“Our research has demonstrated unequivocally that the AP2-G transcription factor protein is essential for flipping the switch that initiates the transformation of malaria parasites in the blood from the asexual stage to the critical sexual stage of their life cycle,” Dr Llinás said.
He and his colleagues believe their discovery is exciting for the future of malaria research. It could spur the development of a sexual-stage vaccine, which would help a person infected with malaria mount an immune response to prevent their parasites from being transmitted to a mosquito, effectively ending the life cycle for that person’s batch of malaria parasites. ![]()

gametocyte stage (blue) and
uninfected red blood cells
Credit: The Llinás lab
Results of 2 new studies suggest that a single regulatory protein acts as a master switch to trigger development of the sexual forms of malaria parasites.
It appears that the protein, AP2-G, is necessary for activating a set of genes that initiate the development of Plasmodium gametocytes, the only forms of the parasite that are infectious to mosquitoes.
This suggests that if researchers can target AP2-G, they can stop sexual parasites from forming.
And if the sexual forms of the parasite never develop in an infected person’s blood, none will enter the mosquito’s gut, and the mosquito will be unable to infect anyone else with malaria.
“Exciting opportunities now lie ahead for finding an effective way to break the chain of malaria transmission by preventing the malaria parasite from completing its full lifecycle,” said Manuel Llinás, PhD, a professor at Pennsylvania State University who was involved in both studies.
The 2 studies, which were published as letters to Nature, had remarkably similar results, despite the fact that the groups worked with 2 different malaria parasites—Plasmodium falciparum and Plasmodium berghei.
In one study, researchers analyzed the whole-genome sequences of 2 P falciparum strains that were unable to produce gametocytes. The only mutated, non-functional gene common to both strains was the AP2-G gene.
In the other study, researchers sequenced P berghei parasites that had lost their ability to make gametocytes. Again, the only common mutated gene in these parasites was AP2-G.
To confirm these observations, both groups of researchers disabled the AP2-G gene in parasites that could generate gametocytes.
As expected, disabling the gene prevented the parasites from producing gametocytes. But the parasites regained their ability to make gametocytes when the mutated gene was repaired.
These results, as well as results of additional experiments, suggest that sexual-stage malaria parasites are produced only when the AP2-G protein is in working order.
“Our research has demonstrated unequivocally that the AP2-G transcription factor protein is essential for flipping the switch that initiates the transformation of malaria parasites in the blood from the asexual stage to the critical sexual stage of their life cycle,” Dr Llinás said.
He and his colleagues believe their discovery is exciting for the future of malaria research. It could spur the development of a sexual-stage vaccine, which would help a person infected with malaria mount an immune response to prevent their parasites from being transmitted to a mosquito, effectively ending the life cycle for that person’s batch of malaria parasites. ![]()

gametocyte stage (blue) and
uninfected red blood cells
Credit: The Llinás lab
Results of 2 new studies suggest that a single regulatory protein acts as a master switch to trigger development of the sexual forms of malaria parasites.
It appears that the protein, AP2-G, is necessary for activating a set of genes that initiate the development of Plasmodium gametocytes, the only forms of the parasite that are infectious to mosquitoes.
This suggests that if researchers can target AP2-G, they can stop sexual parasites from forming.
And if the sexual forms of the parasite never develop in an infected person’s blood, none will enter the mosquito’s gut, and the mosquito will be unable to infect anyone else with malaria.
“Exciting opportunities now lie ahead for finding an effective way to break the chain of malaria transmission by preventing the malaria parasite from completing its full lifecycle,” said Manuel Llinás, PhD, a professor at Pennsylvania State University who was involved in both studies.
The 2 studies, which were published as letters to Nature, had remarkably similar results, despite the fact that the groups worked with 2 different malaria parasites—Plasmodium falciparum and Plasmodium berghei.
In one study, researchers analyzed the whole-genome sequences of 2 P falciparum strains that were unable to produce gametocytes. The only mutated, non-functional gene common to both strains was the AP2-G gene.
In the other study, researchers sequenced P berghei parasites that had lost their ability to make gametocytes. Again, the only common mutated gene in these parasites was AP2-G.
To confirm these observations, both groups of researchers disabled the AP2-G gene in parasites that could generate gametocytes.
As expected, disabling the gene prevented the parasites from producing gametocytes. But the parasites regained their ability to make gametocytes when the mutated gene was repaired.
These results, as well as results of additional experiments, suggest that sexual-stage malaria parasites are produced only when the AP2-G protein is in working order.
“Our research has demonstrated unequivocally that the AP2-G transcription factor protein is essential for flipping the switch that initiates the transformation of malaria parasites in the blood from the asexual stage to the critical sexual stage of their life cycle,” Dr Llinás said.
He and his colleagues believe their discovery is exciting for the future of malaria research. It could spur the development of a sexual-stage vaccine, which would help a person infected with malaria mount an immune response to prevent their parasites from being transmitted to a mosquito, effectively ending the life cycle for that person’s batch of malaria parasites. ![]()
Drug could enhance effects of chemo
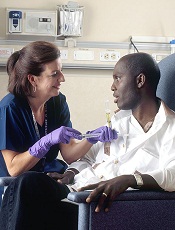
Credit: Rhoda Baer
The drug spironolactone could improve the efficacy of platinum-based chemotherapy by preventing tumor cell repair, according to research published in Chemistry & Biology.
The researchers knew that platinum-based chemotherapy drugs bind to cellular DNA to induce damage.
So they theorized that blocking DNA repair mechanisms would help potentiate chemotherapy by reducing cancer cells’ resistance to treatment.
The team focused their efforts on inhibiting nucleotide excision repair (NER), in which a damaged DNA fragment is replaced with an intact fragment.
Frédéric Coin, PhD, of the Institute of Genetics and Molecular and Cellular Biology in Illkirch, France, and his colleagues screened more than 1200 drugs looking for one that would inhibit NER activity.
And they found that spironolactone—a drug already used to treat fluid retention, high blood pressure, and other conditions—affects NER activity.
Specifically, the team found that, when combined with platinum derivatives, spironolactone significantly increased cytotoxicity in ovarian and colon cancer cells.
As platinum-based chemotherapy is used to treat a range of cancers, similar results might occur in other malignancies as well.
The researchers also noted that, because spironolactone is already in use for other purposes, it doesn’t require a new application for marketing authorization. And its side effects are already known.
The team said this suggests that protocols testing spironolactone in combination with platinum-based chemotherapy could be organized rather quickly. ![]()

Credit: Rhoda Baer
The drug spironolactone could improve the efficacy of platinum-based chemotherapy by preventing tumor cell repair, according to research published in Chemistry & Biology.
The researchers knew that platinum-based chemotherapy drugs bind to cellular DNA to induce damage.
So they theorized that blocking DNA repair mechanisms would help potentiate chemotherapy by reducing cancer cells’ resistance to treatment.
The team focused their efforts on inhibiting nucleotide excision repair (NER), in which a damaged DNA fragment is replaced with an intact fragment.
Frédéric Coin, PhD, of the Institute of Genetics and Molecular and Cellular Biology in Illkirch, France, and his colleagues screened more than 1200 drugs looking for one that would inhibit NER activity.
And they found that spironolactone—a drug already used to treat fluid retention, high blood pressure, and other conditions—affects NER activity.
Specifically, the team found that, when combined with platinum derivatives, spironolactone significantly increased cytotoxicity in ovarian and colon cancer cells.
As platinum-based chemotherapy is used to treat a range of cancers, similar results might occur in other malignancies as well.
The researchers also noted that, because spironolactone is already in use for other purposes, it doesn’t require a new application for marketing authorization. And its side effects are already known.
The team said this suggests that protocols testing spironolactone in combination with platinum-based chemotherapy could be organized rather quickly. ![]()

Credit: Rhoda Baer
The drug spironolactone could improve the efficacy of platinum-based chemotherapy by preventing tumor cell repair, according to research published in Chemistry & Biology.
The researchers knew that platinum-based chemotherapy drugs bind to cellular DNA to induce damage.
So they theorized that blocking DNA repair mechanisms would help potentiate chemotherapy by reducing cancer cells’ resistance to treatment.
The team focused their efforts on inhibiting nucleotide excision repair (NER), in which a damaged DNA fragment is replaced with an intact fragment.
Frédéric Coin, PhD, of the Institute of Genetics and Molecular and Cellular Biology in Illkirch, France, and his colleagues screened more than 1200 drugs looking for one that would inhibit NER activity.
And they found that spironolactone—a drug already used to treat fluid retention, high blood pressure, and other conditions—affects NER activity.
Specifically, the team found that, when combined with platinum derivatives, spironolactone significantly increased cytotoxicity in ovarian and colon cancer cells.
As platinum-based chemotherapy is used to treat a range of cancers, similar results might occur in other malignancies as well.
The researchers also noted that, because spironolactone is already in use for other purposes, it doesn’t require a new application for marketing authorization. And its side effects are already known.
The team said this suggests that protocols testing spironolactone in combination with platinum-based chemotherapy could be organized rather quickly.
Supercomputer accelerates whole-genome analysis

Credit: NIGMS
The time needed to sequence an entire human genome has decreased greatly in recent years, but analyzing the resulting 3 billion base pairs of genetic information from a single genome can take many months.
Now, researchers have found they can accelerate whole-genome analysis using a Cray XE6 supercomputer.
The team found this computer could process many genomes at once and was able to analyze 240 full genomes in a little over 2 days.
The researchers reported these results in Bioinformatics.
The team used Beagle, a Cray XE6 supercomputer located at Argonne National Laboratory in Illinois, in an attempt to analyze multiple genomes concurrently.
Using publicly available software packages and one quarter of its total capacity, the computer was able to align and call variants on 240 whole genomes in approximately 50 hours.
But the computer did not only speed up whole-genome analysis. It also increased the usable sequences per genome.
“Improving analysis through both speed and accuracy reduces the price per genome,” said study author Elizabeth McNally, MD, PhD, of the University of Chicago.
“With this approach, the price for analyzing an entire genome is less than the cost of looking at just a fraction of the genome. New technology promises to bring the costs of sequencing down to around $1000 per genome. Our goal is get the cost of analysis down into that range.”
The findings of this research have immediate medical applications, according to Dr McNally. She noted that she and her colleagues must often sequence genes from an initial patient as well as multiple family members in order to better understand and either treat or prevent a disease.
“We start genetic testing with the patient,” she said. “But when we find a significant mutation, we have to think about testing the whole family to identify individuals at risk.”
Furthermore, the range of testable mutations has greatly increased in recent years.
“In the early days, we would test 1 to 3 genes,” Dr McNally said. “In 2007, we did our first 5-gene panel. Now, we order 50 to 70 genes at a time, which usually gets us an answer. At that point, it can be more useful and less expensive to sequence the whole genome.”
The information from these genomes combined with careful attention to patient and family histories adds to our knowledge about inherited disorders, according to Dr McNally.
“It can refine the classification of these disorders,” she said. “By paying close attention to family members with genes that place them at increased risk, but who do not yet show signs of disease, we can investigate early phases of a disorder.”

Credit: NIGMS
The time needed to sequence an entire human genome has decreased greatly in recent years, but analyzing the resulting 3 billion base pairs of genetic information from a single genome can take many months.
Now, researchers have found they can accelerate whole-genome analysis using a Cray XE6 supercomputer.
The team found this computer could process many genomes at once and was able to analyze 240 full genomes in a little over 2 days.
The researchers reported these results in Bioinformatics.
The team used Beagle, a Cray XE6 supercomputer located at Argonne National Laboratory in Illinois, in an attempt to analyze multiple genomes concurrently.
Using publicly available software packages and one quarter of its total capacity, the computer was able to align and call variants on 240 whole genomes in approximately 50 hours.
But the computer did not only speed up whole-genome analysis. It also increased the usable sequences per genome.
“Improving analysis through both speed and accuracy reduces the price per genome,” said study author Elizabeth McNally, MD, PhD, of the University of Chicago.
“With this approach, the price for analyzing an entire genome is less than the cost of looking at just a fraction of the genome. New technology promises to bring the costs of sequencing down to around $1000 per genome. Our goal is get the cost of analysis down into that range.”
The findings of this research have immediate medical applications, according to Dr McNally. She noted that she and her colleagues must often sequence genes from an initial patient as well as multiple family members in order to better understand and either treat or prevent a disease.
“We start genetic testing with the patient,” she said. “But when we find a significant mutation, we have to think about testing the whole family to identify individuals at risk.”
Furthermore, the range of testable mutations has greatly increased in recent years.
“In the early days, we would test 1 to 3 genes,” Dr McNally said. “In 2007, we did our first 5-gene panel. Now, we order 50 to 70 genes at a time, which usually gets us an answer. At that point, it can be more useful and less expensive to sequence the whole genome.”
The information from these genomes combined with careful attention to patient and family histories adds to our knowledge about inherited disorders, according to Dr McNally.
“It can refine the classification of these disorders,” she said. “By paying close attention to family members with genes that place them at increased risk, but who do not yet show signs of disease, we can investigate early phases of a disorder.”

Credit: NIGMS
The time needed to sequence an entire human genome has decreased greatly in recent years, but analyzing the resulting 3 billion base pairs of genetic information from a single genome can take many months.
Now, researchers have found they can accelerate whole-genome analysis using a Cray XE6 supercomputer.
The team found this computer could process many genomes at once and was able to analyze 240 full genomes in a little over 2 days.
The researchers reported these results in Bioinformatics.
The team used Beagle, a Cray XE6 supercomputer located at Argonne National Laboratory in Illinois, in an attempt to analyze multiple genomes concurrently.
Using publicly available software packages and one quarter of its total capacity, the computer was able to align and call variants on 240 whole genomes in approximately 50 hours.
But the computer did not only speed up whole-genome analysis. It also increased the usable sequences per genome.
“Improving analysis through both speed and accuracy reduces the price per genome,” said study author Elizabeth McNally, MD, PhD, of the University of Chicago.
“With this approach, the price for analyzing an entire genome is less than the cost of looking at just a fraction of the genome. New technology promises to bring the costs of sequencing down to around $1000 per genome. Our goal is get the cost of analysis down into that range.”
The findings of this research have immediate medical applications, according to Dr McNally. She noted that she and her colleagues must often sequence genes from an initial patient as well as multiple family members in order to better understand and either treat or prevent a disease.
“We start genetic testing with the patient,” she said. “But when we find a significant mutation, we have to think about testing the whole family to identify individuals at risk.”
Furthermore, the range of testable mutations has greatly increased in recent years.
“In the early days, we would test 1 to 3 genes,” Dr McNally said. “In 2007, we did our first 5-gene panel. Now, we order 50 to 70 genes at a time, which usually gets us an answer. At that point, it can be more useful and less expensive to sequence the whole genome.”
The information from these genomes combined with careful attention to patient and family histories adds to our knowledge about inherited disorders, according to Dr McNally.
“It can refine the classification of these disorders,” she said. “By paying close attention to family members with genes that place them at increased risk, but who do not yet show signs of disease, we can investigate early phases of a disorder.”
Mutations linked to blood vessel disorders
Two groups of researchers have shown that variants in the gene CECR1, which encodes for ADA2, are associated with blood vessel disorders.
One group found that mutations in CECR1 are associated with an ADA2 deficiency syndrome that is characterized by early onset, recurrent strokes and systemic vasculopathy.
The other group discovered that variants in CECR1 can cause polyarteritis nodosa vasculopathy.
Results of both studies appear in NEJM.
ADA2 deficiency syndrome
In the first study, Qing Zhou, PhD, of the National Human Genome Research Institute (NHGRI), and his colleagues used whole-exome sequencing to gain insight into the aforementioned syndrome, which was characterized by early onset lacunar strokes, systemic vasculopathy, and other symptoms.
The researchers performed whole-exome sequencing in 3 patients with the syndrome and their unaffected parents. In comparing 2 of the patients’ exomes to the exomes of their parents, the researchers located 2 variants in CECR1.
Sequencing a third patient revealed another harmful variant of CECR1, in addition to a small genomic deletion that shuts down the second copy of the gene.
The researchers needed only a sequence reading of that single gene to confirm that 3 other patients were affected by variants in CECR1. In all, these 6 patients were compound heterozygous for 8 mutations in CECR1.
The team discovered that these CECR1 mutations lead to the elimination of ADA2, which produces abnormalities and inflammation in blood vessel walls that ultimately result in the syndrome. So the team called the syndrome deficiency of ADA2 (DADA2).
To gain more insight into DADA2, the researchers induced ADA2 deficiency in a zebrafish model. They found that zebrafish embryos that produce less ADA2 than normal embryos have cerebral bleeds similar to those seen in some of the children with DADA2.
“Our study raises the possibility that the ADA2 pathway may contribute to susceptibility to stroke in the more general population,” said Daniel Kastner, MD, PhD, of NHGRI.
“This genome sequencing study expands what has previously been known about vascular biology. The role of ADA2 in such serious human disease is important and suggests that ADA2 variants may contribute to other, more common illnesses.”
Dr Kastner and his colleagues also sequenced the CECR1 gene in 3 patients from Turkey who had some of the symptoms of DADA2. These patients, who had polyarteritis nodosa (n=2) or small-vessel vasculitis (n=1), were homozygous for the p.Gly47Arg mutation.
ADA2 in polyarteritis nodosa
In the second study, researchers found the same mutation (p.Gly47Arg) in patients that emigrated to Israel from the country of Georgia.
“We now know that this mutation exists in the Middle East and in Pakistani populations and that it is not that uncommon,” said Ivona Aksentijevich, MD, of NHGRI. “This is the first time a single gene has been discovered that is involved in causing a system-wide form of vasculitis.”
To uncover this mutation, Paulina Navon-Elkan, MD, of the Shaare Zedek Medical Center in Jerusalem, and her colleagues sequenced 37 patients who had features of polyarteritis nodosa.
Nineteen patients were of Georgian Jewish ancestry. Sixteen of these patients came from 5 families, and 3 patients were unrelated. An additional 14 unrelated patients were of Turkish ancestry. And 4 German patients came from the same family.
The researchers found that, in all the families, vasculitis was a result of recessive mutations in CECR1.
All of the Georgian Jewish patients were homozygous for the p.Gly47Arg mutation. The German patients were compound heterozygous for Arg169Gln and Pro251Leu mutations. And 1 Turkish patient was compound heterozygous for Gly47Val and Trp264Ser mutations.
The researchers also analyzed serum specimens from the patients and found their ADA2 activity was significantly reduced.
Taking these results together, the team concluded that mutations in CECR1 cause loss of ADA2 function that can result in polyarteritis nodosa vasculopathy with a highly varied clinical expression.

Two groups of researchers have shown that variants in the gene CECR1, which encodes for ADA2, are associated with blood vessel disorders.
One group found that mutations in CECR1 are associated with an ADA2 deficiency syndrome that is characterized by early onset, recurrent strokes and systemic vasculopathy.
The other group discovered that variants in CECR1 can cause polyarteritis nodosa vasculopathy.
Results of both studies appear in NEJM.
ADA2 deficiency syndrome
In the first study, Qing Zhou, PhD, of the National Human Genome Research Institute (NHGRI), and his colleagues used whole-exome sequencing to gain insight into the aforementioned syndrome, which was characterized by early onset lacunar strokes, systemic vasculopathy, and other symptoms.
The researchers performed whole-exome sequencing in 3 patients with the syndrome and their unaffected parents. In comparing 2 of the patients’ exomes to the exomes of their parents, the researchers located 2 variants in CECR1.
Sequencing a third patient revealed another harmful variant of CECR1, in addition to a small genomic deletion that shuts down the second copy of the gene.
The researchers needed only a sequence reading of that single gene to confirm that 3 other patients were affected by variants in CECR1. In all, these 6 patients were compound heterozygous for 8 mutations in CECR1.
The team discovered that these CECR1 mutations lead to the elimination of ADA2, which produces abnormalities and inflammation in blood vessel walls that ultimately result in the syndrome. So the team called the syndrome deficiency of ADA2 (DADA2).
To gain more insight into DADA2, the researchers induced ADA2 deficiency in a zebrafish model. They found that zebrafish embryos that produce less ADA2 than normal embryos have cerebral bleeds similar to those seen in some of the children with DADA2.
“Our study raises the possibility that the ADA2 pathway may contribute to susceptibility to stroke in the more general population,” said Daniel Kastner, MD, PhD, of NHGRI.
“This genome sequencing study expands what has previously been known about vascular biology. The role of ADA2 in such serious human disease is important and suggests that ADA2 variants may contribute to other, more common illnesses.”
Dr Kastner and his colleagues also sequenced the CECR1 gene in 3 patients from Turkey who had some of the symptoms of DADA2. These patients, who had polyarteritis nodosa (n=2) or small-vessel vasculitis (n=1), were homozygous for the p.Gly47Arg mutation.
ADA2 in polyarteritis nodosa
In the second study, researchers found the same mutation (p.Gly47Arg) in patients that emigrated to Israel from the country of Georgia.
“We now know that this mutation exists in the Middle East and in Pakistani populations and that it is not that uncommon,” said Ivona Aksentijevich, MD, of NHGRI. “This is the first time a single gene has been discovered that is involved in causing a system-wide form of vasculitis.”
To uncover this mutation, Paulina Navon-Elkan, MD, of the Shaare Zedek Medical Center in Jerusalem, and her colleagues sequenced 37 patients who had features of polyarteritis nodosa.
Nineteen patients were of Georgian Jewish ancestry. Sixteen of these patients came from 5 families, and 3 patients were unrelated. An additional 14 unrelated patients were of Turkish ancestry. And 4 German patients came from the same family.
The researchers found that, in all the families, vasculitis was a result of recessive mutations in CECR1.
All of the Georgian Jewish patients were homozygous for the p.Gly47Arg mutation. The German patients were compound heterozygous for Arg169Gln and Pro251Leu mutations. And 1 Turkish patient was compound heterozygous for Gly47Val and Trp264Ser mutations.
The researchers also analyzed serum specimens from the patients and found their ADA2 activity was significantly reduced.
Taking these results together, the team concluded that mutations in CECR1 cause loss of ADA2 function that can result in polyarteritis nodosa vasculopathy with a highly varied clinical expression.

Two groups of researchers have shown that variants in the gene CECR1, which encodes for ADA2, are associated with blood vessel disorders.
One group found that mutations in CECR1 are associated with an ADA2 deficiency syndrome that is characterized by early onset, recurrent strokes and systemic vasculopathy.
The other group discovered that variants in CECR1 can cause polyarteritis nodosa vasculopathy.
Results of both studies appear in NEJM.
ADA2 deficiency syndrome
In the first study, Qing Zhou, PhD, of the National Human Genome Research Institute (NHGRI), and his colleagues used whole-exome sequencing to gain insight into the aforementioned syndrome, which was characterized by early onset lacunar strokes, systemic vasculopathy, and other symptoms.
The researchers performed whole-exome sequencing in 3 patients with the syndrome and their unaffected parents. In comparing 2 of the patients’ exomes to the exomes of their parents, the researchers located 2 variants in CECR1.
Sequencing a third patient revealed another harmful variant of CECR1, in addition to a small genomic deletion that shuts down the second copy of the gene.
The researchers needed only a sequence reading of that single gene to confirm that 3 other patients were affected by variants in CECR1. In all, these 6 patients were compound heterozygous for 8 mutations in CECR1.
The team discovered that these CECR1 mutations lead to the elimination of ADA2, which produces abnormalities and inflammation in blood vessel walls that ultimately result in the syndrome. So the team called the syndrome deficiency of ADA2 (DADA2).
To gain more insight into DADA2, the researchers induced ADA2 deficiency in a zebrafish model. They found that zebrafish embryos that produce less ADA2 than normal embryos have cerebral bleeds similar to those seen in some of the children with DADA2.
“Our study raises the possibility that the ADA2 pathway may contribute to susceptibility to stroke in the more general population,” said Daniel Kastner, MD, PhD, of NHGRI.
“This genome sequencing study expands what has previously been known about vascular biology. The role of ADA2 in such serious human disease is important and suggests that ADA2 variants may contribute to other, more common illnesses.”
Dr Kastner and his colleagues also sequenced the CECR1 gene in 3 patients from Turkey who had some of the symptoms of DADA2. These patients, who had polyarteritis nodosa (n=2) or small-vessel vasculitis (n=1), were homozygous for the p.Gly47Arg mutation.
ADA2 in polyarteritis nodosa
In the second study, researchers found the same mutation (p.Gly47Arg) in patients that emigrated to Israel from the country of Georgia.
“We now know that this mutation exists in the Middle East and in Pakistani populations and that it is not that uncommon,” said Ivona Aksentijevich, MD, of NHGRI. “This is the first time a single gene has been discovered that is involved in causing a system-wide form of vasculitis.”
To uncover this mutation, Paulina Navon-Elkan, MD, of the Shaare Zedek Medical Center in Jerusalem, and her colleagues sequenced 37 patients who had features of polyarteritis nodosa.
Nineteen patients were of Georgian Jewish ancestry. Sixteen of these patients came from 5 families, and 3 patients were unrelated. An additional 14 unrelated patients were of Turkish ancestry. And 4 German patients came from the same family.
The researchers found that, in all the families, vasculitis was a result of recessive mutations in CECR1.
All of the Georgian Jewish patients were homozygous for the p.Gly47Arg mutation. The German patients were compound heterozygous for Arg169Gln and Pro251Leu mutations. And 1 Turkish patient was compound heterozygous for Gly47Val and Trp264Ser mutations.
The researchers also analyzed serum specimens from the patients and found their ADA2 activity was significantly reduced.
Taking these results together, the team concluded that mutations in CECR1 cause loss of ADA2 function that can result in polyarteritis nodosa vasculopathy with a highly varied clinical expression.
Study reveals potential target for mucositis, GVHD prevention

Results of preclinical research point to a possible way of preventing mucositis, graft-vs-host disease, and other disorders associated with epithelial permeability.
Investigators created a mouse model of mucositis and discovered that interleukin-1 (IL-1) beta, a protein secreted by the stressed mucosa, played an important role in the condition.
But inhibiting IL-1 beta alleviated mucositis. So the researchers speculated that targeting IL-1 beta might prevent mucositis in humans.
Naama Kanarek, a doctoral student at Hebrew University Hadassah Medical School in Jerusalem, and her colleagues described this research in PNAS.
The investigators began by generating a mouse model deficient in a gene encoding the enzyme beta-TrCP. They chose this enzyme because it’s a major regulator of inflammatory cascades.
The team found that beta-TrCP deletion in the gut caused mucosal DNA damage in the mice, mimicking the effects of chemotherapy and irradiation. Similar to human patients, a severe mucositis reaction occurred in mice that were genetically engineered to be beta-TrCP-deficient.
Tracing the pathological basis of the mouse mucositis revealed that the source of the problem was IL-1 beta. IL-1 beta opened the gut lining, allowing gut bacteria to penetrate and destroy the gut interior.
To confirm this finding, the researchers treated mice with an antibody neutralizing IL-1 beta prior to deleting beta-TrCP. They found this prevented the onset of mucositis.
Therefore, the team has proposed that IL-1 receptor agonists should be tested as mucositis prophylaxis in humans. An example is anakinra (Kineret), which is used to treat chronic inflammatory conditions, such as rheumatoid arthritis and Crohn’s disease.
The investigators believe such treatments might also be used to prevent graft-vs-host disease, burn injuries, head and neck trauma, and other disorders associated with
epithelial permeability.

Results of preclinical research point to a possible way of preventing mucositis, graft-vs-host disease, and other disorders associated with epithelial permeability.
Investigators created a mouse model of mucositis and discovered that interleukin-1 (IL-1) beta, a protein secreted by the stressed mucosa, played an important role in the condition.
But inhibiting IL-1 beta alleviated mucositis. So the researchers speculated that targeting IL-1 beta might prevent mucositis in humans.
Naama Kanarek, a doctoral student at Hebrew University Hadassah Medical School in Jerusalem, and her colleagues described this research in PNAS.
The investigators began by generating a mouse model deficient in a gene encoding the enzyme beta-TrCP. They chose this enzyme because it’s a major regulator of inflammatory cascades.
The team found that beta-TrCP deletion in the gut caused mucosal DNA damage in the mice, mimicking the effects of chemotherapy and irradiation. Similar to human patients, a severe mucositis reaction occurred in mice that were genetically engineered to be beta-TrCP-deficient.
Tracing the pathological basis of the mouse mucositis revealed that the source of the problem was IL-1 beta. IL-1 beta opened the gut lining, allowing gut bacteria to penetrate and destroy the gut interior.
To confirm this finding, the researchers treated mice with an antibody neutralizing IL-1 beta prior to deleting beta-TrCP. They found this prevented the onset of mucositis.
Therefore, the team has proposed that IL-1 receptor agonists should be tested as mucositis prophylaxis in humans. An example is anakinra (Kineret), which is used to treat chronic inflammatory conditions, such as rheumatoid arthritis and Crohn’s disease.
The investigators believe such treatments might also be used to prevent graft-vs-host disease, burn injuries, head and neck trauma, and other disorders associated with
epithelial permeability.

Results of preclinical research point to a possible way of preventing mucositis, graft-vs-host disease, and other disorders associated with epithelial permeability.
Investigators created a mouse model of mucositis and discovered that interleukin-1 (IL-1) beta, a protein secreted by the stressed mucosa, played an important role in the condition.
But inhibiting IL-1 beta alleviated mucositis. So the researchers speculated that targeting IL-1 beta might prevent mucositis in humans.
Naama Kanarek, a doctoral student at Hebrew University Hadassah Medical School in Jerusalem, and her colleagues described this research in PNAS.
The investigators began by generating a mouse model deficient in a gene encoding the enzyme beta-TrCP. They chose this enzyme because it’s a major regulator of inflammatory cascades.
The team found that beta-TrCP deletion in the gut caused mucosal DNA damage in the mice, mimicking the effects of chemotherapy and irradiation. Similar to human patients, a severe mucositis reaction occurred in mice that were genetically engineered to be beta-TrCP-deficient.
Tracing the pathological basis of the mouse mucositis revealed that the source of the problem was IL-1 beta. IL-1 beta opened the gut lining, allowing gut bacteria to penetrate and destroy the gut interior.
To confirm this finding, the researchers treated mice with an antibody neutralizing IL-1 beta prior to deleting beta-TrCP. They found this prevented the onset of mucositis.
Therefore, the team has proposed that IL-1 receptor agonists should be tested as mucositis prophylaxis in humans. An example is anakinra (Kineret), which is used to treat chronic inflammatory conditions, such as rheumatoid arthritis and Crohn’s disease.
The investigators believe such treatments might also be used to prevent graft-vs-host disease, burn injuries, head and neck trauma, and other disorders associated with
epithelial permeability.